Thermus thermophilus L11 methyltransferase, PrmA, is dispensable for growth and preferentially modifies free ribosomal protein L11 prior to ribosome assembly
- PMID: 15317787
- PMCID: PMC516821
- DOI: 10.1128/JB.186.17.5819-5825.2004
Thermus thermophilus L11 methyltransferase, PrmA, is dispensable for growth and preferentially modifies free ribosomal protein L11 prior to ribosome assembly
Abstract
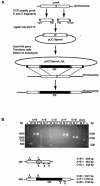
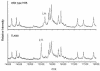
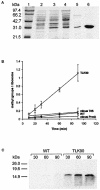
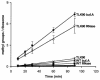
The ribosomal protein L11 in bacteria is posttranslationally trimethylated at multiple amino acid positions by the L11 methyltransferase PrmA, the product of the prmA gene. The role of L11 methylation in ribosome function or assembly has yet to be determined, although the deletion of Escherichia coli prmA has no apparent phenotype. We have constructed a mutant of the extreme thermophile Thermus thermophilus in which the prmA gene has been disrupted with the htk gene encoding a heat-stable kanamycin adenyltransferase. This mutant shows no growth defects, indicating that T. thermophilus PrmA, like its E. coli homolog, is dispensable. Ribosomes prepared from this mutant contain unmethylated L11, as determined by matrix-assisted laser desorption ionization-time of flight mass spectrometry (MALDI-TOF MS), and are effective substrates for in vitro methylation by cloned and purified T. thermophilus PrmA. MALDI-TOF MS also revealed that T. thermophilus L11 contains a total of 12 methyl groups, in contrast to the 9 methyl groups found in E. coli L11. Finally, we found that, as with the E. coli methyltransferase, the ribosomal protein L11 dissociated from ribosomes is a more efficient substrate for in vitro methylation by PrmA than intact 70S ribosomes, suggesting that methylation in vivo occurs on free L11 prior to its incorporation into ribosomes.
Figures
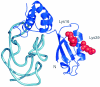
References
-
- Agrawal, R. K., J. Linde, J. Sengupta, K. H. Nierhaus, and J. Frank. 2001. Localization of L11 protein on the ribosome and elucidation of its involvement in EF-G-dependent translocation. J. Mol. Biol. 311:777-787. - PubMed
-
- Alix, J. H. 1992. Extrinsic factors in ribosome assembly, p. 173-184. In K. H. Nierhaus, A. R. Subramanian, V. A. Erdmann, F. Franceschi, and B. Wittmann-Liebold (ed.), The translational apparatus. Plenum Publishing, New York, N.Y.
-
- Allen, P. N., and H. F. Noller. 1991. A single base substitution in 16S ribosomal RNA suppresses streptomycin dependence and increases the frequency of translational errors. Cell 66:141-148. - PubMed
-
- Bujnicki, J. M. 2000. Sequence, structural, and evolutionary analysis of prokaryotic ribosomal protein L11 methyltransferases. Acta Microbiol. Pol. 49:19-28. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

