Transcript profiling in the chl1-5 mutant of Arabidopsis reveals a role of the nitrate transporter NRT1.1 in the regulation of another nitrate transporter, NRT2.1
- PMID: 15319483
- PMCID: PMC520944
- DOI: 10.1105/tpc.104.024380
Transcript profiling in the chl1-5 mutant of Arabidopsis reveals a role of the nitrate transporter NRT1.1 in the regulation of another nitrate transporter, NRT2.1
Abstract
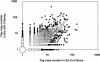
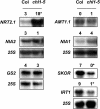
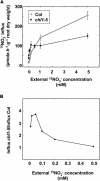
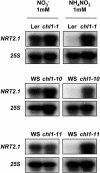
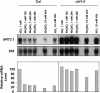
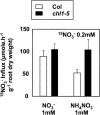
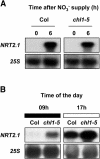
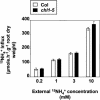
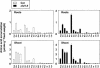
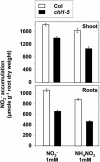
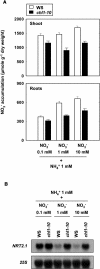
Arabidopsis thaliana mutants deficient for the NRT1.1 NO(3)(-) transporter display complex phenotypes, including lowered NO(3)(-) uptake, altered development of nascent organs, and reduced stomatal opening. To obtain further insight at the molecular level on the multiple physiological functions of NRT1.1, we performed large-scale transcript profiling by serial analysis of gene expression in the roots of the chl1-5 deletion mutant of NRT1.1 and of the Columbia wild type. Several hundred genes were differentially expressed between the two genotypes, when plants were grown on NH(4)NO(3) as N source. Among these genes, the N satiety-repressed NRT2.1 gene, encoding a major component of the root high-affinity NO(3)(-) transport system (HATS), was found to be strongly derepressed in the chl1-5 mutant (as well as in other NRT1.1 mutants). This was associated with a marked stimulation of the NO(3)(-) HATS activity in the mutant, suggesting adaptive response to a possible N limitation resulting from NRT1.1 mutation. However, derepression of NRT2.1 in NH(4)NO(3)-fed chl1-5 plants could not be attributed to lowered production of N metabolites. Rather, the results show that normal regulation of NRT2.1 expression is strongly altered in the chl1-5 mutant, where this gene is no more repressible by high N provision to the plant. This indicates that NRT1.1 plays an unexpected but important role in the regulation of both NRT2.1 expression and NO(3)(-) HATS activity. Overexpression of NRT2.1 was also found in wild-type plants supplied with 1 mM NH(4)(+) plus 0.1 mM NO(3)(-), a situation where NRT1.1 is likely to mediate very low NO(3)(-) transport. Thus, we suggest that it is the lack of NRT1.1 activity, rather than the absence of this transporter, that derepresses NRT2.1 expression in the presence of NH(4)(+). Two hypotheses are discussed to explain these results: (1) NRT2.1 is upregulated by a NO(3)(-) demand signaling, indirectly triggered by lack of NRT1.1-mediated uptake, which overrides feedback repression by N metabolites, and (2) NRT1.1 plays a more direct signaling role, and its transport activity generates an unknown signal required for NRT2.1 repression by N metabolites. Both mechanisms would warrant that either NRT1.1 or NRT2.1 ensure significant NO(3)(-) uptake in the presence of NH(4)(+) in the external medium, which is crucial to prevent the detrimental effects of pure NH(4)(+) nutrition.
Figures
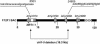

Similar articles
-
Regulation of the high-affinity NO3- uptake system by NRT1.1-mediated NO3- demand signaling in Arabidopsis.Plant Physiol. 2006 Nov;142(3):1075-86. doi: 10.1104/pp.106.087510. Epub 2006 Sep 22. Plant Physiol. 2006. PMID: 16998085 Free PMC article.
-
Characterization of the signalling pathways involved in the repression of root nitrate uptake by nitrate in Arabidopsis thaliana.J Exp Bot. 2023 Aug 3;74(14):4244-4258. doi: 10.1093/jxb/erad149. J Exp Bot. 2023. PMID: 37185665
-
A reevaluation of the contribution of NRT1.1 to nitrate uptake in Arabidopsis under low-nitrate supply.FEBS Lett. 2019 Aug;593(15):2051-2059. doi: 10.1002/1873-3468.13473. Epub 2019 Jun 13. FEBS Lett. 2019. PMID: 31172512
-
Nitrate transporters and peptide transporters.FEBS Lett. 2007 May 25;581(12):2290-300. doi: 10.1016/j.febslet.2007.04.047. Epub 2007 Apr 26. FEBS Lett. 2007. PMID: 17481610 Review.
-
The regulation of nitrate and ammonium transport systems in plants.J Exp Bot. 2002 Apr;53(370):855-64. doi: 10.1093/jexbot/53.370.855. J Exp Bot. 2002. PMID: 11912228 Review.
Cited by
-
Enhancement of nitrogen use efficiency through agronomic and molecular based approaches in cotton.Front Plant Sci. 2022 Sep 27;13:994306. doi: 10.3389/fpls.2022.994306. eCollection 2022. Front Plant Sci. 2022. PMID: 36237509 Free PMC article. Review.
-
Gametophytic selection in Arabidopsis thaliana supports the selective model of intron length reduction.PLoS Genet. 2005 Aug;1(2):e13. doi: 10.1371/journal.pgen.0010013. Epub 2005 Aug 5. PLoS Genet. 2005. PMID: 16110339 Free PMC article.
-
A system biology approach highlights a hormonal enhancer effect on regulation of genes in a nitrate responsive "biomodule".BMC Syst Biol. 2009 Jun 6;3:59. doi: 10.1186/1752-0509-3-59. BMC Syst Biol. 2009. PMID: 19500399 Free PMC article.
-
Increase in tomato locule number is controlled by two single-nucleotide polymorphisms located near WUSCHEL.Plant Physiol. 2011 Aug;156(4):2244-54. doi: 10.1104/pp.111.173997. Epub 2011 Jun 14. Plant Physiol. 2011. PMID: 21673133 Free PMC article.
-
Overexpression of a High-Affinity Nitrate Transporter OsNRT2.1 Increases Yield and Manganese Accumulation in Rice Under Alternating Wet and Dry Condition.Front Plant Sci. 2018 Aug 15;9:1192. doi: 10.3389/fpls.2018.01192. eCollection 2018. Front Plant Sci. 2018. PMID: 30158947 Free PMC article.
References
-
- Clarkson, D.T., Gojon, A., Saker, L.R., Wiersema, P.K., Purves, J.V., Tillard, P., Arnold, G.M., Paams, A.J.M., Waalburg, W., and Stulen, I. (1996). Nitrate and ammonium influxes in soybean (Glycine max) roots: Direct comparison of 13N and 15N tracing. Plant Cell Environ. 19, 859–868.
-
- Crawford, N.M., and Glass, A.D.M. (1998). Molecular and physiological aspects of nitrate uptake in plants. Trends Plant Sci. 3, 389–395.
-
- Delhon, P., Gojon, A., Tillard, P., and Passama, L. (1995). Diurnal regulation of NO3- uptake in soybean plants. I. Changes in NO3- influx, efflux, and N utilization in the plant during the day/night cycle. J. Exp. Bot. 46, 1585–1594.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

