Proteus mirabilis ZapA metalloprotease degrades a broad spectrum of substrates, including antimicrobial peptides
- PMID: 15322010
- PMCID: PMC517467
- DOI: 10.1128/IAI.72.9.5159-5167.2004
Proteus mirabilis ZapA metalloprotease degrades a broad spectrum of substrates, including antimicrobial peptides
Abstract
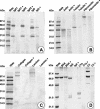
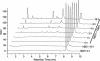
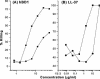
The 54-kDa extracellular metalloprotease ZapA is an important virulence factor of uropathogenic Proteus mirabilis. While ZapA has the ability to degrade host immunoglobulins (Igs), the dramatic attenuation of virulence in ZapA mutants suggests that this enzyme may have a broader spectrum of activity. This hypothesis was tested by in vitro assays with purified ZapA and an array of purified protein or peptide substrates. The data reveal that many proteins found in the urinary tract are substrates of ZapA proteolysis, including complement (C1q and C3), cell matrix (collagen, fibronectin, and laminin), and cytoskeletal proteins (actin and tubulin). Proteolysis of IgA and IgG was significantly enhanced by conditions that denatured the Igs. It was discovered that the antimicrobial peptides human beta-defensin 1 (hBD1) and LL-37 are readily cleaved by the enzyme. To the best of our knowledge, this is the first report of a bacterial protease capable of cleaving hBD1, a component of the human renal tubule innate immune response. Proteolysis of hBD1 resulted in ca. six peptides, while proteolysis of LL-37 resulted in at least nine products. Matrix-assisted laser desorption ionization-time of flight mass spectrometry analysis of the molecular masses of the reaction products indicated that ZapA preferred no distinct peptide bond. The antimicrobial activity of hBD1 and LL-37 was significantly reduced following ZapA treatment, suggesting that proteolysis results in inactivation of these peptides. The data suggest that a function of ZapA during urinary tract infections is the proteolysis of antimicrobial peptides associated with the innate immune response.
Figures




References
-
- Aneas, M. A., F. C. Portaro, I. Lebrun, L. Juliano, M. S. Palma, and B. L. Fernandes. 2001. ZapA, a possible virulence factor from Proteus mirabilis exhibits broad protease substrate specificity. Braz. J. Med. Biol. Res. 34:1397-1403. - PubMed
-
- Bensch, K. W., M. Raida, H. J. Magert, P. Schulz-Knappe, and W. G. Forssmann. 1995. hBD-1: a novel beta-defensin from human plasma. FEBS Lett. 368:331-335. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

