Activation-induced cytidine deaminase (AID) can target both DNA strands when the DNA is supercoiled
- PMID: 15328407
- PMCID: PMC516507
- DOI: 10.1073/pnas.0404974101
Activation-induced cytidine deaminase (AID) can target both DNA strands when the DNA is supercoiled
Abstract
The activation-induced cytidine deaminase (AID) is required for somatic hypermutation (SHM) and class-switch recombination of Ig genes. It has been shown that in vitro, AID protein deaminates C in single-stranded DNA or the coding-strand DNA that is being transcribed but not in double-stranded DNA. However, in vivo, both DNA strands are mutated equally during SHM. We show that AID efficiently deaminates C on both DNA strands of a supercoiled plasmid, acting preferentially on SHM hotspot motifs. However, this DNA is not targeted by AID when it is relaxed after treatment with topoisomerase I, and thus, supercoiling plays a crucial role for AID targeting to this DNA. Most of the mutations are in negatively supercoiled regions, suggesting a mechanism of AID targeting in vivo. During transcription the DNA sequences upstream of the elongating RNA polymerase are negatively supercoiled, and this transient change in DNA topology may allow AID to access both DNA strands.
Copyright 2004 The National Academy of Sciencs of the USA
Figures
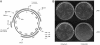

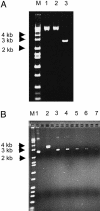

References
-
- Muramatsu, M., Sankaranand, V., Anant, S., Sugai, M., Kinoshita, K., Davidson, N. & Honjo, T. (1999) J. Biol. Chem. 274, 18470–18476. - PubMed
-
- Muramatsu, M., Kinoshita, K., Fagarasan, S., Yamada, S., Shinkai, Y. & Honjo, T. (2000) Cell 102, 553–563. - PubMed
-
- Revy, P., Muto, T., Levy, Y., Geissman, F., Plebani, A., Sanal, O., Catalan, N., Forveille, M., Dufourcq-Lagelouse, R., Gennery, A., et al. (2000) Cell 102, 565–575. - PubMed
-
- Diaz, M. & Storb, U. (2003) DNA Repair 2, 623–627. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

