Crystal structure of Ski8p, a WD-repeat protein with dual roles in mRNA metabolism and meiotic recombination
- PMID: 15340168
- PMCID: PMC2001155
- DOI: 10.1110/ps.04856504
Crystal structure of Ski8p, a WD-repeat protein with dual roles in mRNA metabolism and meiotic recombination
Abstract
Ski8p is a WD-repeat protein with an essential role for the Ski complex assembly in an exosome-dependent 3'-to-5' mRNA decay. In addition, Ski8p is involved in meiotic recombination by interacting with Spo11p protein. We have determined the crystal structure of Ski8p from Saccharomyces cerevisiae at 2.2 A resolution. The structure reveals that Ski8p folds into a seven-bladed beta propeller. Mapping sequence conservation and hydrophobicities of amino acids on the molecular surface of Ski8p reveals a prominent site on the top surface of the beta propeller, which is most likely involved in mediating interactions of Ski8p with Ski3p and Spo11p. Mutagenesis combined with yeast two-hybrid and GST pull-down assays identified the top surface of the beta propeller as being required for Ski8p binding to Ski3p and Spo11p. The functional implications for Ski8p function in both mRNA decay and meiotic recombination are discussed.
Figures
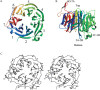





Similar articles
-
The structure of Ski8p, a protein regulating mRNA degradation: Implications for WD protein structure.Protein Sci. 2004 Jun;13(6):1557-65. doi: 10.1110/ps.04704704. Protein Sci. 2004. PMID: 15152089 Free PMC article.
-
The yeast antiviral proteins Ski2p, Ski3p, and Ski8p exist as a complex in vivo.RNA. 2000 Mar;6(3):449-57. doi: 10.1017/s1355838200991787. RNA. 2000. PMID: 10744028 Free PMC article.
-
Identification of residues in yeast Spo11p critical for meiotic DNA double-strand break formation.Mol Cell Biol. 2002 Feb;22(4):1106-15. doi: 10.1128/MCB.22.4.1106-1115.2002. Mol Cell Biol. 2002. PMID: 11809802 Free PMC article.
-
Mechanism and control of meiotic recombination initiation.Curr Top Dev Biol. 2001;52:1-53. doi: 10.1016/s0070-2153(01)52008-6. Curr Top Dev Biol. 2001. PMID: 11529427 Review.
-
Functions of the yeast meiotic recombination genes, MRE11 and MRE2.Adv Biophys. 1995;31:67-76. doi: 10.1016/0065-227x(95)99383-z. Adv Biophys. 1995. PMID: 7625279 Review.
Cited by
-
Decay of mRNAs targeted by RISC requires XRN1, the Ski complex, and the exosome.RNA. 2005 Apr;11(4):459-69. doi: 10.1261/rna.7231505. Epub 2005 Feb 9. RNA. 2005. PMID: 15703439 Free PMC article.
-
The carboxy-terminal domain of Erb1 is a seven-bladed ß-propeller that binds RNA.PLoS One. 2015 Apr 16;10(4):e0123463. doi: 10.1371/journal.pone.0123463. eCollection 2015. PLoS One. 2015. PMID: 25880847 Free PMC article.
-
Ge-1 is a central component of the mammalian cytoplasmic mRNA processing body.RNA. 2005 Dec;11(12):1795-802. doi: 10.1261/rna.2142405. RNA. 2005. PMID: 16314453 Free PMC article.
-
Functional interactions among members of the meiotic initiation complex in fission yeast.Curr Genet. 2010 Jun;56(3):237-49. doi: 10.1007/s00294-010-0296-0. Epub 2010 Apr 3. Curr Genet. 2010. PMID: 20364342
-
Identifying the hotspots on the top faces of WD40-repeat proteins from their primary sequences by β-bulges and DHSW tetrads.PLoS One. 2012;7(8):e43005. doi: 10.1371/journal.pone.0043005. Epub 2012 Aug 15. PLoS One. 2012. PMID: 22916195 Free PMC article.
References
-
- Aloy, P., Bottcher, B., Ceulemans, H., Leutwein, C., Mellwig, C., Fischer, S., Gavin, A.C., Bork, P., Superti-Furga, G., Serrano, L., et al. 2004. Structure-based assembly of protein complexes in yeast. Science 303 2026–2029. - PubMed
-
- Arora, C., Kee, K., Maleki, S., and Keeney, S. 2004. Antiviral protein Ski8 is a direct partner of Spo11 in meiotic DNA break formation, independent of its cytoplasmic role in RNA metabolism. Mol. Cell 13 549–559. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

