Display of bacterial lipase on the Escherichia coli cell surface by using FadL as an anchoring motif and use of the enzyme in enantioselective biocatalysis
- PMID: 15345384
- PMCID: PMC520891
- DOI: 10.1128/AEM.70.9.5074-5080.2004
Display of bacterial lipase on the Escherichia coli cell surface by using FadL as an anchoring motif and use of the enzyme in enantioselective biocatalysis
Abstract
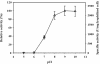
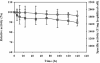
We have developed a novel cell surface display system by employing FadL as an anchoring motif, which is an outer membrane protein involved in long-chain fatty acid transport in Escherichia coli. A thermostable Bacillus sp. strain TG43 lipase (44.5 kDa) could be successfully displayed on the cell surface of E. coli in an active form by C-terminal deletion-fusion of lipase at the ninth external loop of FadL. The localization of the truncated FadL-lipase fusion protein on the cell surface was confirmed by confocal microscopy and Western blot analysis. Lipase activity was mainly detected with whole cells, but not with the culture supernatant, suggesting that cell lysis was not a problem. The activity of cell surface-displayed lipase was examined at different temperatures and pHs and was found to be the highest at 50 degrees C and pH 9 to 10. Cell surface-displayed lipase was quite stable, even at 60 and 70 degrees C, and retained over 90% of the full activity after incubation at 50 degrees C for a week. As a potential application, cell surface-displayed lipase was used as a whole-cell catalyst for kinetic resolution of racemic methyl mandelate. In 36 h of reaction, (S)-mandelic acid could be produced with the enantiomeric excess of 99% and the enantiomeric ratio of 292, which are remarkably higher than values obtained with crude lipase or cross-linked lipase crystal. These results suggest that FadL may be a useful anchoring motif for displaying enzymes on the cell surface of E. coli for whole-cell biocatalysis.
Figures




References
-
- Ahmed, S. N., R. J. Kazlauskas, A. H. Morinville, P. Grochulski, J. D. Schrag, and M. W. Cygler. 1994. Enantioselectivity of Candida rugosa lipase toward carboxylic acids: a predictive rule from substrate mapping and X-ray crystallography. Biocatalysis 9:209-225.
-
- Bell, P. J. L., H. Nevalainen, H. W. Morgan, and P. L. Bergquist. 1999. Rapid cloning of thermoalkalophilic lipases from Bacillus spp. using PCR. Biotechnol. Lett. 21:1003-1006.
-
- Benhar, I. 2001. Biotechnological applications of phage and cell display. Biotechnol. Adv. 19:1-33. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Research Materials

