Regulation of the N-acyl homoserine lactone-dependent quorum-sensing system in rhizosphere Pseudomonas putida WCS358 and cross-talk with the stationary-phase RpoS sigma factor and the global regulator GacA
- PMID: 15345437
- PMCID: PMC520884
- DOI: 10.1128/AEM.70.9.5493-5502.2004
Regulation of the N-acyl homoserine lactone-dependent quorum-sensing system in rhizosphere Pseudomonas putida WCS358 and cross-talk with the stationary-phase RpoS sigma factor and the global regulator GacA
Abstract
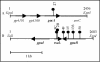
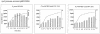
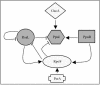
Quorum sensing is a cell population-density dependent regulatory system which in gram-negative bacteria often involves the production and detection of N-acyl homoserine lactones (AHLs). Some Pseudomonas putida strains have been reported to produce AHLs, and one quorum-sensing locus has been identified. However, it appears that the majority of strains do not produce AHLs. In this study we report the identification and regulation of the AHL-dependent system of rhizosphere P. putida WCS358. This system is identical to the recently identified system of P. putida strain IsoF and very similar to the las system of Pseudomonas aeruginosa. It is composed of three genes, the luxI family member ppuI, the putative repressor rsaL, and the luxR family member ppuR. A genomic ppuR::Tn5 mutant of strain WCS358 was identified by its inability to produce AHLs when it was cross-streaked in close proximity to an AHL biosensor, whereas an rsaL::Tn5 genomic mutant was identified by its ability to overproduce AHL molecules. Using transcriptional promoter fusions, we studied expression profiles of the rsaL, ppuI, and ppuR promoters in various genetic backgrounds. At the onset of the stationary phase, the autoinducer synthase ppuI gene expression is under positive regulation by PpuR-AHL and under negative regulation by RsaL, indicating that the molecules could be in competition for binding at the ppuI promoter. In genomic rsaL::Tn5 mutants ppuI expression and production of AHL levels increased dramatically; however, both processes were still under growth phase regulation, indicating that RsaL is not involved in repressing AHL production at low cell densities. The roles of the global response regulator GacA and the stationary-phase sigma factor RpoS in the regulation of the AHL system at the onset of the stationary phase were also investigated. The P. putida WCS358 gacA gene was cloned and inactivated in the genome. It was determined that the three global regulatory systems are closely linked, with quorum sensing and RpoS regulating each other and GacA positively regulating ppuI expression. Studies of the regulation of AHL quorum-sensing systems have lagged behind other studies and are important for understanding how these systems are integrated into the overall growth phase and metabolic status of the cells.
Figures






Similar articles
-
The Pseudomonas putida Lon protease is involved in N-acyl homoserine lactone quorum sensing regulation.BMC Microbiol. 2007 Jul 26;7:71. doi: 10.1186/1471-2180-7-71. BMC Microbiol. 2007. PMID: 17655747 Free PMC article.
-
The ppuI-rsaL-ppuR quorum-sensing system regulates biofilm formation of Pseudomonas putida PCL1445 by controlling biosynthesis of the cyclic lipopeptides putisolvins I and II.J Bacteriol. 2006 Apr;188(8):2898-906. doi: 10.1128/JB.188.8.2898-2906.2006. J Bacteriol. 2006. PMID: 16585751 Free PMC article.
-
Identification and characterization of an N-acylhomoserine lactone-dependent quorum-sensing system in Pseudomonas putida strain IsoF.Appl Environ Microbiol. 2002 Dec;68(12):6371-82. doi: 10.1128/AEM.68.12.6371-6382.2002. Appl Environ Microbiol. 2002. PMID: 12450862 Free PMC article.
-
An evolving perspective on the Pseudomonas aeruginosa orphan quorum sensing regulator QscR.Front Cell Infect Microbiol. 2014 Oct 28;4:152. doi: 10.3389/fcimb.2014.00152. eCollection 2014. Front Cell Infect Microbiol. 2014. PMID: 25389523 Free PMC article. Review.
-
Control of rpoS transcription in Escherichia coli and Pseudomonas: why so different?Mol Microbiol. 2003 Jul;49(1):1-9. doi: 10.1046/j.1365-2958.2003.03547.x. Mol Microbiol. 2003. PMID: 12823806 Review.
Cited by
-
Chromosomal Arrangement of AHL-Driven Quorum Sensing Circuits in Pseudomonas.ISRN Microbiol. 2012 Feb 29;2012:484176. doi: 10.5402/2012/484176. Print 2012. ISRN Microbiol. 2012. PMID: 23724324 Free PMC article.
-
Functional characterization of the quorum sensing regulator RsaL in the plant-beneficial strain Pseudomonas putida WCS358.Appl Environ Microbiol. 2012 Feb;78(3):726-34. doi: 10.1128/AEM.06442-11. Epub 2011 Nov 23. Appl Environ Microbiol. 2012. PMID: 22113916 Free PMC article.
-
PpoR is a conserved unpaired LuxR solo of Pseudomonas putida which binds N-acyl homoserine lactones.BMC Microbiol. 2009 Jun 17;9:125. doi: 10.1186/1471-2180-9-125. BMC Microbiol. 2009. PMID: 19534812 Free PMC article.
-
The Systematic Investigation of the Quorum Sensing System of the Biocontrol Strain Pseudomonas chlororaphis subsp. aurantiaca PB-St2 Unveils aurI to Be a Biosynthetic Origin for 3-Oxo-Homoserine Lactones.PLoS One. 2016 Nov 18;11(11):e0167002. doi: 10.1371/journal.pone.0167002. eCollection 2016. PLoS One. 2016. PMID: 27861617 Free PMC article.
-
Two-component system GacS/GacA, a global response regulator of bacterial physiological behaviors.Eng Microbiol. 2022 Oct 5;3(1):100051. doi: 10.1016/j.engmic.2022.100051. eCollection 2023 Mar. Eng Microbiol. 2022. PMID: 39628522 Free PMC article. Review.
References
-
- Bertani, I., M. Kojic, and V. Venturi. 2001. Regulation of the p-hydroxybenzoic acid hydroxylase gene (pobA) in plant-growth-promoting Pseudomonas putida WCS358. Microbiology 147:1611-1620. - PubMed
-
- Better, M., B. Lewis, D. Corbin, G. Ditta, and D. R. Helinski. 1983. Structural relationships among Rhizobium meliloti symbiotic promoters. Cell 35:479-485. - PubMed
-
- Birnboim, H. C. 1983. A rapid alkaline extraction method for the isolation of plasmid DNA. Methods Enzymol. 100:243-255. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
LinkOut - more resources
Full Text Sources

