Alkaline anaerobic respiration: isolation and characterization of a novel alkaliphilic and metal-reducing bacterium
- PMID: 15345448
- PMCID: PMC520920
- DOI: 10.1128/AEM.70.9.5595-5602.2004
Alkaline anaerobic respiration: isolation and characterization of a novel alkaliphilic and metal-reducing bacterium
Abstract
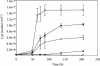
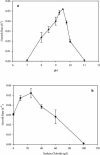
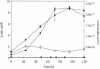
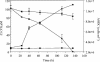
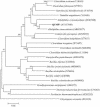
Iron-reducing enrichments were obtained from leachate ponds at the U.S. Borax Company in Boron, Calif. Based on partial small-subunit (SSU) rRNA gene sequences (approximately 500 nucleotides), six isolates shared 98.9% nucleotide identity. As a representative, the isolate QYMF was selected for further analysis. QYMF could be grown with Fe(III)-citrate, Fe(III)-EDTA, Co(III)-EDTA, or Cr(VI) as electron acceptors, and yeast extract and lactate could serve as electron donors. Growth during iron reduction occurred over the pH range of 7.5 to 11.0 (optimum, pH 9.5), a sodium chloride range of 0 to 80 g/liter (optimum, 20 g/liter), and a temperature range of 4 to 45 degrees C (optimum, approximately 35 degrees C), and iron precipitates were formed. QYMF was a strict anaerobe that could be grown in the presence of borax, and the cells were straight rods that produced endospores. Sodium chloride and yeast extract stimulated growth. Phylogenetic analysis of the SSU rRNA gene indicated that the bacterium was a low-G+C gram-positive microorganism and had 96 and 92% nucleotide identity with Alkaliphilus transvaalensis and Alkaliphilus crotonatoxidans, respectively. The major phospholipid fatty acids were 14:1, 16:1omega7c, and 16:0, which were different from those of other alkaliphiles but similar to those of reported iron-reducing bacteria. The results demonstrated that the isolate might represent a novel metal-reducing alkaliphilic species. The name Alkaliphilus metalliredigens sp. nov. is proposed. The isolation and activity of metal-reducing bacteria from borax-contaminated leachate ponds suggest that bioremediation of metal-contaminated alkaline environments may be feasible and have implications for alkaline anaerobic respiration.
Figures



References
-
- Blair, B. G., and K. L. Anderson. 1998. Comparison of staining techniques for scanning electron microscopic detection of ultrastructural protuberances on cellulolytic bacteria. Biotech. Histochem. 73:107-113. - PubMed
-
- Bligh, E. G., and W. J. Dyer. 1959. A rapid method of total lipid extraction and purification. Can. J. Biochem. Physiol. 37:911-917. - PubMed
-
- Cao, X., X. Liu, and X. Dong. 2003. Alkaliphilus crotonatoxidans sp. nov., a strictly anaerobic, crotonate-dismutating bacterium isolated from a methanogenic environment. Int. J. Syst. E vol. Microbiol. 53:971-975. - PubMed
-
- Duckworth, A. W., W. D. Grant, B. E. Jones, and R. van Steenbergen. 1996. Phylogenetic diversity of soda lake alkaliphiles. FEMS Microbiol. Ecol. 19:181-191.
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

