Cloning and characterization of the bile salt hydrolase genes (bsh) from Bifidobacterium bifidum strains
- PMID: 15345449
- PMCID: PMC520925
- DOI: 10.1128/AEM.70.9.5603-5612.2004
Cloning and characterization of the bile salt hydrolase genes (bsh) from Bifidobacterium bifidum strains
Abstract
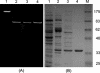
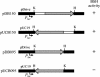
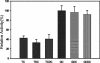
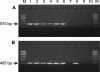
Biochemical characterization of the purified bile salt hydrolase (BSH) from Bifidobacterium bifidum ATCC 11863 revealed some distinct characteristics not observed in other species of Bifidobacterium. The bsh gene was cloned from B. bifidum, and the DNA flanking the bsh gene was sequenced. Comparison of the deduced amino acid sequence of the cloned gene with previously known sequences revealed high homology with BSH enzymes from several microorganisms and penicillin V amidase (PVA) of Bacillus sphaericus. The proposed active sites of PVA were highly conserved, including that of the Cys-1 residue. The importance of the SH group in the N-terminal cysteine was confirmed by substitution of Cys with chemically and structurally similar residues, Ser or Thr, both of which resulted in an inactive enzyme. The transcriptional start point of the bsh gene has been determined by primer extension analysis. Unlike Bifidobacterium longum bsh, B. bifidum bsh was transcribed as a monocistronic unit, which was confirmed by Northern blot analysis. PCR amplification with the type-specific primer set revealed the high level of sequence homology in their bsh genes within the species of B. bifidum.
Figures




Similar articles
-
Bile salt hydrolase of Bifidobacterium longum-biochemical and genetic characterization.Appl Environ Microbiol. 2000 Jun;66(6):2502-12. doi: 10.1128/AEM.66.6.2502-2512.2000. Appl Environ Microbiol. 2000. PMID: 10831430 Free PMC article.
-
Cloning and characterization of a bile salt hydrolase (bsh) from Bifidobacterium adolescentis.Biotechnol Lett. 2005 Jun;27(12):817-22. doi: 10.1007/s10529-005-6717-3. Biotechnol Lett. 2005. PMID: 16086241
-
Genetic analysis of a bile salt hydrolase in Bifidobacterium animalis subsp. lactis KL612.J Appl Microbiol. 2008 Sep;105(3):778-90. doi: 10.1111/j.1365-2672.2008.03825.x. Epub 2008 Apr 25. J Appl Microbiol. 2008. PMID: 18444998
-
Genus- and species-specific PCR primers for the detection and identification of bifidobacteria.Curr Issues Intest Microbiol. 2003 Sep;4(2):61-9. Curr Issues Intest Microbiol. 2003. PMID: 14503690 Review.
-
Functional and Phylogenetic Diversity of BSH and PVA Enzymes.Microorganisms. 2021 Mar 31;9(4):732. doi: 10.3390/microorganisms9040732. Microorganisms. 2021. PMID: 33807488 Free PMC article. Review.
Cited by
-
A selective gut bacterial bile salt hydrolase alters host metabolism.Elife. 2018 Jul 17;7:e37182. doi: 10.7554/eLife.37182. Elife. 2018. PMID: 30014852 Free PMC article.
-
Developing a procedure to extract chenodeoxycholic acid and synthesize ursodeoxycholic acid from pig by-products.Heliyon. 2023 Jul 17;9(7):e18313. doi: 10.1016/j.heliyon.2023.e18313. eCollection 2023 Jul. Heliyon. 2023. PMID: 37519734 Free PMC article.
-
Functional analysis of four bile salt hydrolase and penicillin acylase family members in Lactobacillus plantarum WCFS1.Appl Environ Microbiol. 2008 Aug;74(15):4719-26. doi: 10.1128/AEM.00137-08. Epub 2008 Jun 6. Appl Environ Microbiol. 2008. PMID: 18539794 Free PMC article.
-
A bile salt hydrolase of Brucella abortus contributes to the establishment of a successful infection through the oral route in mice.Infect Immun. 2007 Jan;75(1):299-305. doi: 10.1128/IAI.00952-06. Epub 2006 Nov 6. Infect Immun. 2007. PMID: 17088355 Free PMC article.
-
Systematic identification of secondary bile acid production genes in global microbiome.mSystems. 2025 Jan 21;10(1):e0081724. doi: 10.1128/msystems.00817-24. Epub 2024 Dec 17. mSystems. 2025. PMID: 39688414 Free PMC article.
References
-
- Asahara, T., K. Shimizu, K. Nomoto, M. Watanuki, and R. Tanaka. 2001. Antibacterial effect of fermented milk containing Bifidobacterium breve, Bifidobacterium bifidum, and Lactobacillus acidophilus against indigenous Escherichia coli infection in mice. Microb. Ecol. Health Dis. 13:16-24.
-
- Bahar, R. J., and S. Andrew. 1999. Bile acid transport. Gastroenterol. Clin. N. Am. 28:27-58. - PubMed
-
- Ballongue, J. 1998. Bifidobacteria and probiotic action, p. 519-587. In S. Salminen and A. Von Wright (ed.), Lactic acid bacteria. Microbiology and functional aspects. Marcel Dekker, Inc., New York, N.Y.
-
- Batta, A. K., G. Salen, R. Arora, S. Shefer, M. Batta, and A. Person. 1990. Side chain conjugation prevents bacterial 7-dehydroxylation of bile acids. J. Biol. Chem. 265:10925-10928. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

