Microscopic Kinetics of DNA Translocation through synthetic nanopores
- PMID: 15345583
- PMCID: PMC1304610
- DOI: 10.1529/biophysj.104.042960
Microscopic Kinetics of DNA Translocation through synthetic nanopores
Abstract
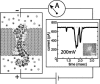
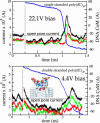
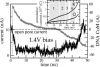
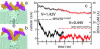
We have previously demonstrated that a nanometer-diameter pore in a nanometer-thick metal-oxide-semiconductor-compatible membrane can be used as a molecular sensor for detecting DNA. The prospects for using this type of device for sequencing DNA are avidly being pursued. The key attribute of the sensor is the electric field-induced (voltage-driven) translocation of the DNA molecule in an electrolytic solution across the membrane through the nanopore. To complement ongoing experimental studies developing such pores and measuring signals in response to the presence of DNA, we conducted molecular dynamics simulations of DNA translocation through the nanopore. A typical simulated system included a patch of a silicon nitride membrane dividing water solution of potassium chloride into two compartments connected by the nanopore. External electrical fields induced capturing of the DNA molecules by the pore from the solution and subsequent translocation. Molecular dynamics simulations suggest that 20-basepair segments of double-stranded DNA can transit a nanopore of 2.2 x 2.6 nm(2) cross section in a few microseconds at typical electrical fields. Hydrophobic interactions between DNA bases and the pore surface can slow down translocation of single-stranded DNA and might favor unzipping of double-stranded DNA inside the pore. DNA occluding the pore mouth blocks the electrolytic current through the pore; these current blockades were found to have the same magnitude as the blockade observed when DNA transits the pore. The feasibility of using molecular dynamics simulations to relate the level of the blocked ionic current to the sequence of DNA was investigated.
Figures





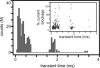

References
-
- Allen, M. P., and D. J. Tildesley. 1987. Computer Simulation of Liquids. Oxford University Press, New York.
-
- Ambjörnsson, T., S. P. Apell, Z. Konkoli, E. A. D. Marzio, and J. J. Kasianowicz. 2002. Charged polymer membrane translocation. Biophys. J. 117:4063–4073.
-
- Balaeff, A., C. R. Koudella, L. Mahadevan, and K. Schulten. 2004. Modeling DNA loops using continuum and statistical mechanics. Philos. Trans. R. Soc. Lond. A. In press. - PubMed
-
- Batcho, P. F., D. A. Case, and T. Schlick. 2001. Optimized particle-mesh Ewald/multiple-time step integration for molecular dynamics simulations. J. Chem. Phys. 115:4003–4018.
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

