Comparative structural modeling and inference of conserved protein classes in Drosophila seminal fluid
- PMID: 15345744
- PMCID: PMC518759
- DOI: 10.1073/pnas.0405579101
Comparative structural modeling and inference of conserved protein classes in Drosophila seminal fluid
Abstract
The constituents of seminal fluid are a complex mixture of proteins and other molecules, most of whose functions have yet to be determined and many of which are rapidly evolving. As a step in elucidating the roles of these proteins and exposing potential functional similarities hidden by their rapid evolution, we performed comparative structural modeling on 28 of 52 predicted seminal proteins produced in the Drosophila melanogaster male accessory gland. Each model was characterized by defining residues likely to be important for structure and function. Comparisons of known protein structures with predicted accessory gland proteins (Acps) revealed similarities undetectable by primary sequence alignments. The structures predict that Acps fall into several categories: regulators of proteolysis, lipid modifiers, immunity/protection, sperm-binding proteins, and peptide hormones. The comparative structural modeling approach indicates that major functional classes of mammalian and Drosophila seminal fluid proteins are conserved, despite differences in reproductive strategies. This is particularly striking in the face of the rapid protein sequence evolution that characterizes many reproductive proteins, including Drosophila and mammalian seminal proteins.
Figures

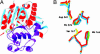

References
-
- Wolfner, M. F. (2002) Heredity 88, 85–93. - PubMed
-
- Wolfner, M. F., Harada, H. A., Bertram, M. J., Stelick, T. J., Kraus, K. W., Kalb, J. M., Lung, Y. O., Neubaum, D. M., Park, M. & Tram, U. (1997) Insect Biochem. Mol. Biol. 27, 825–834. - PubMed
-
- Monsma, S. A., Harada, H. A. & Wolfner, M. F. (1990) Dev. Biol. 142, 465–475. - PubMed
-
- Chen, P. S., Stumm-Zollinger, E., Aigaki, T., Balmer, J., Bienz, M. & Bohlen, P. (1988) Cell 54, 291–298. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

