Real-time fluorescence PCR assays for detection and characterization of heat-labile I and heat-stable I enterotoxin genes from enterotoxigenic Escherichia coli
- PMID: 15364995
- PMCID: PMC516355
- DOI: 10.1128/JCM.42.9.4092-4100.2004
Real-time fluorescence PCR assays for detection and characterization of heat-labile I and heat-stable I enterotoxin genes from enterotoxigenic Escherichia coli
Abstract
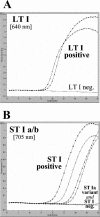
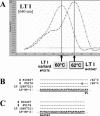
To facilitate the diagnosis of enterotoxigenic Escherichia coli (ETEC) infections in humans, we developed and evaluated real-time fluorescence PCR assays for the Roche LightCycler (LC) against the enterotoxin genes commonly present in strains associated with human illness. Separate LC-PCR assays with identical cycling conditions were designed for the type I heat-labile enterotoxin (LT I) and the type I heat-stable enterotoxin (ST I) genes, using the LC hybridization probe format. A duplex assay for ST I with two sets of amplification primers and three hybridization probes was required to detect the major nucleotide sequence variants of ST I, ST Ia and ST Ib. LC-PCR findings from the testing of 161 E. coli isolates of human origin (138 ETEC and 23 non-ETEC) were compared with those obtained by block cycler PCR analysis. The sensitivities and specificities of the LC-PCR assays were each 100% for the LT I and ST I genes. The LC-PCR and block cycler PCR assays were also compared for their abilities to detect LT I and ST I genes in spiked stool specimens with different methods of sample preparation. Findings from these experiments revealed that the limits of detection for the LC-PCR assays were the same or substantially lower than those observed for the block cycler PCR assay. Melting curve analysis of the amplified LT I and ST I genes revealed sequence variation within each gene, which for the ST I genes correlated with the presence of ST Ia and ST Ib. The rapidity, sensitivity, and specificity of the LC-PCR assays make them attractive alternatives to block cycler PCR assays for the detection and characterization of ETEC.
Figures

References
-
- Black, R.E. 1986. The epidemiology of cholera and enterotoxigenic Escherichia coli diarrheal disease, p. 23-32. In J. Holmgren, A. Lindberg, and R. Mollby (ed.), Development of vaccine and drug against diarrhea. Eleventh Nobel Conference, Stockholm. Studentlitteratur, Lund, Sweden.
-
- Centers for Disease Control and Prevention. 1994. Foodborne outbreaks of enterotoxigenic Escherichia coli—Rhode Island and New Hampshire. Morb. Mortal. Wkly. Rep. 43:81-89. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

