Structure and mechanistic analysis of the anti-human immunodeficiency virus type 1 antibody 2F5 in complex with its gp41 epitope
- PMID: 15367639
- PMCID: PMC516390
- DOI: 10.1128/JVI.78.19.10724-10737.2004
Structure and mechanistic analysis of the anti-human immunodeficiency virus type 1 antibody 2F5 in complex with its gp41 epitope
Abstract
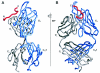
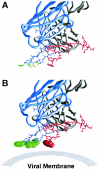
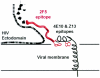
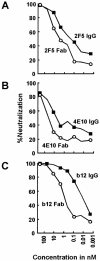
The membrane-proximal region of the ectodomain of the gp41 envelope glycoprotein of human immunodeficiency virus type 1 (HIV-1) is the target of three of the five broadly neutralizing anti-HIV-1 antibodies thus far isolated. We have determined crystal structures of the antigen-binding fragment for one of these antibodies, 2F5, in complex with 7-mer, 11-mer, and 17-mer peptides of the gp41 membrane-proximal region, at 2.0-, 2.1-, and 2.2-A resolutions, respectively. The structures reveal an extended gp41 conformation, which stretches over 30 A in length. Contacts are made with five complementarity-determining regions of the antibody as well as with nonpolymorphic regions. Only one exclusive charged face of the gp41 epitope is bound by 2F5, while the nonbound face, which is hydrophobic, may be hidden due to occlusion by other portions of the ectodomain. The structures reveal that the 2F5 antibody is uniquely built to bind to an epitope that is proximal to a membrane surface and in a manner mostly unaffected by large-scale steric hindrance. Biochemical studies with proteoliposomes confirm the importance of lipid membrane and hydrophobic context in the binding of 2F5 as well as in the binding of 4E10, another broadly neutralizing antibody that recognizes the membrane-proximal region of gp41. Based on these structural and biochemical results, immunization strategies for eliciting 2F5- and 4E10-like broadly neutralizing anti-HIV-1 antibodies are proposed.
Figures





References
-
- Babcock, G. J., T. Mirzabekov, W. Wojtowicz, and J. Sodroski. 2001. Ligand binding characteristics of CXCR4 incorporated into paramagnetic proteoliposomes. J. Biol. Chem. 276:38433-38440. - PubMed
-
- Barbato, G., E. Bianchi, P. Ingallinella, W. H. Hurni, M. D. Miller, G. Ciliberto, R. Cortese, R. Bazzo, J. W. Shiver, and A. Pessi. 2003. Structural analysis of the epitope of the anti-HIV antibody 2F5 sheds light into its mechanism of neutralization and HIV fusion. J. Mol. Biol. 330:1101-1115. - PubMed
-
- Biron, Z., S. Khare, A. O. Samson, Y. Hayek, F. Naider, and J. Anglister. 2002. A monomeric 3(10)-helix is formed in water by a 13-residue peptide representing the neutralizing determinant of HIV-1 on gp41. Biochemistry 41:12687-12696. - PubMed
-
- Brunger, A. T., P. D. Adams, G. M. Clore, W. L. DeLano, P. Gros, R. W. Grosse-Kunstleve, J. S. Jiang, J. Kuszewski, M. Nilges, N. S. Pannu, R. J. Read, L. M. Rice, T. Simonson, and G. L. Warren. 1998. Crystallography & NMR system: a new software suite for macromolecular structure determination. Acta Crystallogr. D 54:905-921. - PubMed
-
- Bryson, S., A. Cunningham, J. Ho, R. C. Hynes, D. E. Isenman, B. H. Barber, R. Kunert, H. Katinger, M. Klein, and E. F. Pai. 2001. Cross-neutralizing human monoclonal anti-HIV-1 antibody 2F5: preparation and crystallographic analysis of the free and epitope-complexed forms of its Fab′ fragment. Protein Peptide Lett. 8:413-418.
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

