Temporal modulation of an autoprotease is crucial for replication and pathogenicity of an RNA virus
- PMID: 15367643
- PMCID: PMC516412
- DOI: 10.1128/JVI.78.19.10765-10775.2004
Temporal modulation of an autoprotease is crucial for replication and pathogenicity of an RNA virus
Abstract
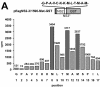
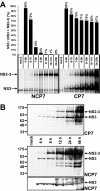
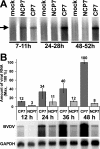
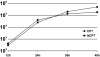
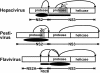
Pestiviruses belong to the family Flaviviridae, and their genome is a single-stranded RNA of positive polarity encoding one large polyprotein which is further processed into mature proteins. Noncytopathogenic (noncp) strains of the pestivirus bovine viral diarrhea virus (BVDV) can establish persistent infection. In persistently infected animals, noncp BVDVs occasionally acquire mutations in viral nonstructural protein 2 (NS2) that give rise to cytopathogenic (cp) BVDV variants, and, eventually, lead to the onset of lethal disease. A molecular marker of cp BVDV infection is a high-level expression of the replicative NS3 protease/helicase that together with NS2 is derived from NS2-3. Here, we present evidence for NS2-3 autoprocessing by a newly identified cysteine protease in NS2 that is distantly related to the NS2-3 autoprotease of hepatitis C and GB viruses. The vital role of this autoprotease in BVDV infection was established, implying an essential function for NS3 in pestiviral RNA replication which cannot be supplied by its NS2-3 precursor. Accordingly, and contrary to a current paradigm, we detected almost complete cleavage of NS2-3 in noncp BVDV at early hours of infection. At 6 to 9 h postinfection, NS2-3 autoprocessing diminished to barely detectable levels for noncp BVDV but decreased only moderately for cp BVDV. Viral RNA synthesis rates strictly correlated with different NS3 levels in noncp and cp BVDV-infected cells, implicating the NS2 autoprotease in RNA replication control. The biotype-specific modulation of NS2-3 autoprocessing indicates a crucial role of the NS2 autoprotease in the pathogenicity of BVDV.
Figures




Similar articles
-
CRISPR/Cas9-Mediated Knockout of DNAJC14 Verifies This Chaperone as a Pivotal Host Factor for RNA Replication of Pestiviruses.J Virol. 2019 Feb 19;93(5):e01714-18. doi: 10.1128/JVI.01714-18. Print 2019 Mar 1. J Virol. 2019. PMID: 30518653 Free PMC article.
-
Establishment and characterization of cytopathogenic and noncytopathogenic pestivirus replicons.J Virol. 1999 Nov;73(11):9422-32. doi: 10.1128/JVI.73.11.9422-9432.1999. J Virol. 1999. PMID: 10516051 Free PMC article.
-
Bovine viral diarrhea virus strain Oregon: a novel mechanism for processing of NS2-3 based on point mutations.J Virol. 1998 May;72(5):4127-38. doi: 10.1128/JVI.72.5.4127-4138.1998. J Virol. 1998. PMID: 9557702 Free PMC article.
-
Pathogenesis of mucosal disease, a deadly disease of cattle caused by a pestivirus.Clin Diagn Virol. 1998 Jul 15;10(2-3):121-7. doi: 10.1016/s0928-0197(98)00037-3. Clin Diagn Virol. 1998. PMID: 9741637 Review.
-
Non-structural proteins of bovine viral diarrhea virus.Virus Genes. 2022 Dec;58(6):491-500. doi: 10.1007/s11262-022-01914-8. Epub 2022 May 25. Virus Genes. 2022. PMID: 35614328 Free PMC article. Review.
Cited by
-
A double deletion prevents replication of the pestivirus bovine viral diarrhea virus in the placenta of pregnant heifers.PLoS Pathog. 2021 Dec 8;17(12):e1010107. doi: 10.1371/journal.ppat.1010107. eCollection 2021 Dec. PLoS Pathog. 2021. PMID: 34879119 Free PMC article.
-
Intracellular restriction of a productive noncytopathic coronavirus infection.J Virol. 2008 Jan;82(1):451-60. doi: 10.1128/JVI.01251-07. Epub 2007 Oct 24. J Virol. 2008. PMID: 17959675 Free PMC article.
-
The NS5A protein of bovine viral diarrhea virus contains an essential zinc-binding site similar to that of the hepatitis C virus NS5A protein.J Virol. 2006 Aug;80(15):7450-8. doi: 10.1128/JVI.00358-06. J Virol. 2006. PMID: 16840325 Free PMC article.
-
Inhibition of sphingosine kinase by bovine viral diarrhea virus NS3 is crucial for efficient viral replication and cytopathogenesis.J Biol Chem. 2009 May 15;284(20):13648-13659. doi: 10.1074/jbc.M807498200. Epub 2009 Mar 17. J Biol Chem. 2009. PMID: 19293152 Free PMC article.
-
Stability of the resistance to the thiosemicarbazone derived from 5,6-dimethoxy-1-indanone, a non-nucleoside polymerase inhibitor of bovine viral diarrhea virus.PLoS One. 2014 Jun 20;9(6):e100528. doi: 10.1371/journal.pone.0100528. eCollection 2014. PLoS One. 2014. PMID: 24950191 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

