Connecting two-component regulatory systems by a protein that protects a response regulator from dephosphorylation by its cognate sensor
- PMID: 15371344
- PMCID: PMC517523
- DOI: 10.1101/gad.1230804
Connecting two-component regulatory systems by a protein that protects a response regulator from dephosphorylation by its cognate sensor
Abstract
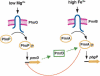
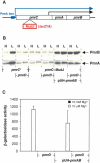
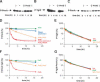
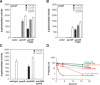
A fundamental question in signal transduction is how an organism integrates multiple signals into a cellular response. Here we report the mechanism by which the Salmonella PmrA/PmrB two-component system responds to the signal controlling the PhoP/PhoQ two-component system. We establish that the PhoP-activated PmrD protein binds to the phosphorylated form of the response regulator PmrA, preventing both its intrinsic dephosphorylation and that promoted by its cognate sensor kinase PmrB. This results in PmrA-mediated transcription because phosphorylated PmrA exhibits higher affinity for its target promoters than unphosphorylated PmrA. A PmrD-independent form of the PmrA protein was resistant to PmrB-catalyzed dephosphorylation and promoted transcription of PmrA-activated genes in the absence of inducing signals. This is the first example of a protein that enables a two-component system to respond to the signal governing a different two-component system by protecting the phosphorylated form of a response regulator.
Figures


Similar articles
-
A small protein that mediates the activation of a two-component system by another two-component system.EMBO J. 2000 Apr 17;19(8):1861-72. doi: 10.1093/emboj/19.8.1861. EMBO J. 2000. PMID: 10775270 Free PMC article.
-
Closing the loop: the PmrA/PmrB two-component system negatively controls expression of its posttranscriptional activator PmrD.Proc Natl Acad Sci U S A. 2003 Apr 15;100(8):4706-11. doi: 10.1073/pnas.0836837100. Epub 2003 Apr 3. Proc Natl Acad Sci U S A. 2003. PMID: 12676988 Free PMC article.
-
Transcriptional regulation of the 4-amino-4-deoxy-L-arabinose biosynthetic genes in Yersinia pestis.J Biol Chem. 2005 Apr 15;280(15):14765-72. doi: 10.1074/jbc.M413900200. Epub 2005 Feb 14. J Biol Chem. 2005. PMID: 15710615
-
The biology of the PmrA/PmrB two-component system: the major regulator of lipopolysaccharide modifications.Annu Rev Microbiol. 2013;67:83-112. doi: 10.1146/annurev-micro-092412-155751. Epub 2013 Jun 17. Annu Rev Microbiol. 2013. PMID: 23799815 Free PMC article. Review.
-
The pleiotropic two-component regulatory system PhoP-PhoQ.J Bacteriol. 2001 Mar;183(6):1835-42. doi: 10.1128/JB.183.6.1835-1842.2001. J Bacteriol. 2001. PMID: 11222580 Free PMC article. Review. No abstract available.
Cited by
-
Dam methylation participates in the regulation of PmrA/PmrB and RcsC/RcsD/RcsB two component regulatory systems in Salmonella enterica serovar Enteritidis.PLoS One. 2013;8(2):e56474. doi: 10.1371/journal.pone.0056474. Epub 2013 Feb 13. PLoS One. 2013. PMID: 23418573 Free PMC article.
-
Feedback Control of Two-Component Regulatory Systems.Annu Rev Microbiol. 2016 Sep 8;70:103-24. doi: 10.1146/annurev-micro-102215-095331. Annu Rev Microbiol. 2016. PMID: 27607549 Free PMC article. Review.
-
A connector of two-component regulatory systems promotes signal amplification and persistence of expression.Proc Natl Acad Sci U S A. 2007 Jul 17;104(29):12063-8. doi: 10.1073/pnas.0704462104. Epub 2007 Jul 5. Proc Natl Acad Sci U S A. 2007. PMID: 17615238 Free PMC article.
-
Gene expression kinetics governs stimulus-specific decoration of the Salmonella outer membrane.Sci Signal. 2018 May 8;11(529):eaar7921. doi: 10.1126/scisignal.aar7921. Sci Signal. 2018. PMID: 29739882 Free PMC article.
-
Phosphoethanolamine modification of lipid A in colistin-resistant variants of Acinetobacter baumannii mediated by the pmrAB two-component regulatory system.Antimicrob Agents Chemother. 2011 Jul;55(7):3370-9. doi: 10.1128/AAC.00079-11. Epub 2011 May 16. Antimicrob Agents Chemother. 2011. PMID: 21576434 Free PMC article.
References
-
- Atkinson M.R., Kamberov, E.S., Weiss, R.L., and Ninfa, A.J. 1994. Reversible uridylylation of the Escherichia coli PII signal transduction protein regulates its ability to stimulate the dephosphorylation of the transcription factor nitrogen regulator I (NRI or NtrC). J. Biol. Chem. 269: 28288-28293. - PubMed
-
- Bijlsma J.J. and Groisman, E.A. 2003. Making informed decisions: Regulatory interactions between two-component systems. Trends Microbiol. 11: 359-366. - PubMed
-
- Burkholder W.F., Kurtser, I., and Grossman, A.D. 2001. Replication initiation proteins regulate a developmental checkpoint in Bacillus subtilis. Cell 104: 269-279. - PubMed
-
- Cherepanov P.P. and Wackernagel, W. 1995. Gene disruption in Escherichia coli: TcR and KmR cassettes with the option of Flp-catalyzed excision of the antibiotic-resistance determinant. Gene 158: 9-14. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
