Variable Surface Glycoprotein RoTat 1.2 PCR as a specific diagnostic tool for the detection of Trypanosoma evansi infections
- PMID: 15377385
- PMCID: PMC521498
- DOI: 10.1186/1475-9292-3-3
Variable Surface Glycoprotein RoTat 1.2 PCR as a specific diagnostic tool for the detection of Trypanosoma evansi infections
Abstract
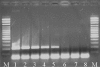
BACKGROUND: Based on the recently sequenced gene coding for the Trypanosoma evansi (T. evansi) RoTat 1.2 Variable Surface Glycoprotein (VSG), a primer pair was designed targeting the DNA region lacking homology to other known VSG genes. A total of 39 different trypanosome stocks were tested using the RoTat 1.2 based Polymerase Chain Reaction (PCR). RESULTS: This PCR yielded a 205 bp product in all T. evansi and in seven out of nine T. equiperdum strains tested. This product was not detected in the DNA from T. b. brucei, T. b. gambiense, T. b. rhodesiense, T. congolense, T. vivax and T. theileri parasites. The Rotat 1.2 PCR detects as few as 10 trypanosomes per reaction with purified DNA from blood samples, i.e. 50 trypanosomes/ml. CONCLUSION: PCR amplification of the RoTat 1.2 VSG gene is a specific marker for T. evansi strains, except T. evansi type B, and is especially useful in dyskinetoplastic strains where kDNA based markers may fail to amplify. Furthermore, our data support previous suggestions that some T. evansi stocks have been previously misclassified as T. equiperdum.
Figures

Similar articles
-
PCR amplification of RoTat 1.2 VSG gene in Trypanosoma evansi isolates in Kenya.Vet Parasitol. 2004 Feb 26;120(1-2):23-33. doi: 10.1016/j.vetpar.2003.12.007. Vet Parasitol. 2004. PMID: 15019140
-
The detection of non-RoTat 1.2 Trypanosoma evansi.Exp Parasitol. 2005 May;110(1):30-8. doi: 10.1016/j.exppara.2005.01.001. Exp Parasitol. 2005. PMID: 15804376
-
Genotyping of Trypanosoma brucei evansi in Egyptian camels: detection of a different non-RoTat 1.2 Trypanosoma brucei evansi in Egyptian camels.Trop Anim Health Prod. 2023 Jul 28;55(4):279. doi: 10.1007/s11250-023-03673-6. Trop Anim Health Prod. 2023. PMID: 37505344 Free PMC article.
-
The expression of RoTat 1.2 variable surface glycoprotein (VSG) in Trypanosoma evansi and T. equiperdum.Vet Parasitol. 2003 Oct 20;116(3):209-16. doi: 10.1016/s0304-4017(02)00359-x. Vet Parasitol. 2003. PMID: 14559163
-
Expression of RoTat 1.2 cross-reactive variable antigen type in Trypanosoma evansi and T. equiperdum.Ann N Y Acad Sci. 2002 Oct;969:174-9. doi: 10.1111/j.1749-6632.2002.tb04373.x. Ann N Y Acad Sci. 2002. PMID: 12381586
Cited by
-
Development of a loop-mediated isothermal amplification assay based on RoTat1.2 gene for detection of Trypanosoma evansi in domesticated animals.Parasitol Res. 2021 May;120(5):1873-1882. doi: 10.1007/s00436-021-07118-7. Epub 2021 Mar 13. Parasitol Res. 2021. PMID: 33712930
-
Dourine: a neglected disease of equids.Trop Anim Health Prod. 2017 Jun;49(5):887-897. doi: 10.1007/s11250-017-1280-1. Epub 2017 Apr 24. Trop Anim Health Prod. 2017. PMID: 28439783 Free PMC article. Review.
-
Population sub-structuring among Trypanosoma evansi stocks.Parasitol Res. 2007 Oct;101(5):1215-24. doi: 10.1007/s00436-007-0603-y. Epub 2007 Jun 22. Parasitol Res. 2007. PMID: 17587054
-
Epidemiology of Trypanosoma evansi and Trypanosoma vivax in domestic animals from selected districts of Tigray and Afar regions, Northern Ethiopia.Parasit Vectors. 2015 Apr 9;8:212. doi: 10.1186/s13071-015-0818-1. Parasit Vectors. 2015. PMID: 25889702 Free PMC article.
-
A review on the diagnosis of animal trypanosomoses.Parasit Vectors. 2022 Feb 19;15(1):64. doi: 10.1186/s13071-022-05190-1. Parasit Vectors. 2022. PMID: 35183235 Free PMC article. Review.
References
-
- Hoare CA. The trypanosomes of mammals. Oxford, Blackwell Scientific Publications; 1972. pp. 1–749.
-
- Stephen LE. In: Trypanosomiasis A veterinary perspective. StephenLE, editor. Oxford, Pergamon Press; 1986.
-
- Nantulya VM. Trypanosomiasis in domestic animals: the problems of diagnosis. Rev Sci Tech. 1990;9:357–367. - PubMed
-
- Moser DR, Cook GA, Ochs DE, Bailey CP, McKane MR, Donelson JE. Detection of Trypanosoma congolense and Trypanosoma brucei subspecies by DNA amplification using the polymerase chain reaction. Parasitology. 1989;99:57–66. - PubMed
-
- Wuyts N, Chokesajjawatee N, Panyim S. A simplified and highly sensitivity detection of Trypanosoma evansi by DNA amplification. Southeast Asian J Trop Med Public Health. 1994;25:266–271. - PubMed
LinkOut - more resources
Full Text Sources

