The post-replication repair RAD18 and RAD6 genes are involved in the prevention of spontaneous mutations caused by 7,8-dihydro-8-oxoguanine in Saccharomyces cerevisiae
- PMID: 15388802
- PMCID: PMC521648
- DOI: 10.1093/nar/gkh831
The post-replication repair RAD18 and RAD6 genes are involved in the prevention of spontaneous mutations caused by 7,8-dihydro-8-oxoguanine in Saccharomyces cerevisiae
Abstract
7,8-dihydro-8-oxoguanine (8-oxoG) is an abundant and mutagenic lesion produced in DNA exposed to free radicals and reactive oxygen species. In Saccharomyces cerevisiae, the OGG1 gene encodes the 8-oxoG DNA N-glycosylase/AP lyase (Ogg1), which is the functional homologue of the bacterial Fpg. Ogg1-deficient strains are spontaneous mutators that accumulate GC to TA transversions due to unrepaired 8-oxoG in DNA. In yeast, DNA mismatch repair (MMR) and translesion synthesis (TLS) by DNA polymerase eta also play a role in the prevention of the mutagenic effect of 8-oxoG. In the present study, we show the RAD18 and RAD6 genes that are required to initiate post-replication repair (PRR) are also involved in the prevention of mutations by 8-oxoG. Consistently, a synergistic increase in spontaneous CanR and Lys+ mutation rates is observed in the absence of Rad6 or Rad18 proteins in ogg1 mutant strains. Spectra of CaR mutations in ogg1 rad18 and ogg1 rad6 double mutants show a strong bias in the favor of GC to TA transversions, which are 137- and 189-fold higher than in the wild-type, respectively. The results also show that Poleta (RAD30 gene product) plays a critical role on the prevention of mutations at 8-oxoG, whereas Polzeta (REV3 gene product) does not. Our current model suggests that the Rad6-Rad18 complex targets Poleta at DNA gaps that result from the MMR-mediated excision of adenine mispaired with 8-oxoG, allowing error-free dCMP incorporation opposite to this lesion.
Figures
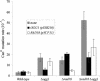

Similar articles
-
PCNA monoubiquitylation and DNA polymerase eta ubiquitin-binding domain are required to prevent 8-oxoguanine-induced mutagenesis in Saccharomyces cerevisiae.Nucleic Acids Res. 2009 May;37(8):2549-59. doi: 10.1093/nar/gkp105. Epub 2009 Mar 5. Nucleic Acids Res. 2009. PMID: 19264809 Free PMC article.
-
Repair of 8-oxo-7,8-dihydroguanine in prokaryotic and eukaryotic cells: Properties and biological roles of the Fpg and OGG1 DNA N-glycosylases.Free Radic Biol Med. 2017 Jun;107:179-201. doi: 10.1016/j.freeradbiomed.2016.11.042. Epub 2016 Nov 27. Free Radic Biol Med. 2017. PMID: 27903453 Review.
-
Specificities of the Saccharomyces cerevisiae rad6, rad18, and rad52 mutators exhibit different degrees of dependence on the REV3 gene product, a putative nonessential DNA polymerase.Genetics. 1995 Jun;140(2):443-56. doi: 10.1093/genetics/140.2.443. Genetics. 1995. PMID: 7498727 Free PMC article.
-
Repair of 8-oxoguanine in Saccharomyces cerevisiae: interplay of DNA repair and replication mechanisms.Free Radic Biol Med. 2002 Jun 15;32(12):1244-53. doi: 10.1016/s0891-5849(02)00822-5. Free Radic Biol Med. 2002. PMID: 12057762 Review.
-
Cloning and expression in Escherichia coli of the OGG1 gene of Saccharomyces cerevisiae, which codes for a DNA glycosylase that excises 7,8-dihydro-8-oxoguanine and 2,6-diamino-4-hydroxy-5-N-methylformamidopyrimidine.Proc Natl Acad Sci U S A. 1996 May 28;93(11):5197-202. doi: 10.1073/pnas.93.11.5197. Proc Natl Acad Sci U S A. 1996. PMID: 8643552 Free PMC article.
Cited by
-
DNA repair mechanisms and the bypass of DNA damage in Saccharomyces cerevisiae.Genetics. 2013 Apr;193(4):1025-64. doi: 10.1534/genetics.112.145219. Genetics. 2013. PMID: 23547164 Free PMC article. Review.
-
In vivo bypass of 8-oxodG.PLoS Genet. 2013;9(8):e1003682. doi: 10.1371/journal.pgen.1003682. Epub 2013 Aug 1. PLoS Genet. 2013. PMID: 23935538 Free PMC article.
-
PCNA monoubiquitylation and DNA polymerase eta ubiquitin-binding domain are required to prevent 8-oxoguanine-induced mutagenesis in Saccharomyces cerevisiae.Nucleic Acids Res. 2009 May;37(8):2549-59. doi: 10.1093/nar/gkp105. Epub 2009 Mar 5. Nucleic Acids Res. 2009. PMID: 19264809 Free PMC article.
-
Yeast-based assays for the functional characterization of cancer-associated variants of human DNA repair genes.Microb Cell. 2020 May 18;7(7):162-174. doi: 10.15698/mic2020.07.721. Microb Cell. 2020. PMID: 32656256 Free PMC article. Review.
-
Role of PCNA-dependent stimulation of 3'-phosphodiesterase and 3'-5' exonuclease activities of human Ape2 in repair of oxidative DNA damage.Nucleic Acids Res. 2009 Jul;37(13):4247-55. doi: 10.1093/nar/gkp357. Epub 2009 May 13. Nucleic Acids Res. 2009. PMID: 19443450 Free PMC article.
References
-
- Lindahl T. (1993) Instability and decay of the primary structure of DNA. Nature, 362, 709–715. - PubMed
-
- Lindahl T. and Wood,R.D. (1999) Quality control by DNA repair. Science, 286, 1897–1905. - PubMed
-
- Hoeijmakers J.H.J. (2001) Genome maintenance mechanisms for preventing cancer. Nature, 411, 366–374. - PubMed
-
- Boiteux S. and Guillet,M. (2004) Abasic sites in DNA: repair and biological consequences in Saccharomyces cerevisiae. DNA Repair, 3, 1–12. - PubMed
-
- Marnett L.J. (2000) Oxyradicals and DNA damage. Carcinogenesis, 21, 361–370. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

