Selection and evolution of NTP-specific aptamers
- PMID: 15452272
- PMCID: PMC521651
- DOI: 10.1093/nar/gkh835
Selection and evolution of NTP-specific aptamers
Abstract
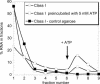
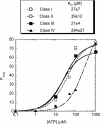
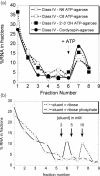
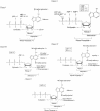
ATP occupies a central position in biology, for it is both an elementary building block of RNA and the most widely used cofactor in all living organisms. For this reason, it has been a recurrent target for in vitro molecular evolution techniques. The exploration of ATP-binding motifs constitutes both an important step in investigating the plausibility of the 'RNA world' hypothesis and a central starting point for the development of new enzymes. To date, only two RNA motifs that bind ATP have been characterized. The first one is targeted to the adenosine moiety, while the second one recognizes the 'Hoogsteen' face of the base. To isolate aptamers that bind ATP in different orientations, we selected RNAs on an affinity resin that presents ATP in three different orientations. We obtained five new motifs that were characterized and subsequently submitted to a secondary selection protocol designed to isolate aptamers specific for cordycepin. Interestingly, all the ATP-binding motifs selected specifically recognize the sugar-phosphate backbone region of the nucleotides. Three of the aptamers show some selectivity for adenine derivatives, while the remainder recognize any of the four nucleotides with similar efficiency. The characteristics of these aptamers are discussed along with implications for in vitro molecular evolution.
Figures








Similar articles
-
Boron-containing aptamers to ATP.Nucleic Acids Res. 2002 Mar 15;30(6):1401-7. doi: 10.1093/nar/30.6.1401. Nucleic Acids Res. 2002. PMID: 11884639 Free PMC article.
-
RNA aptamers to the adenosine moiety of S-adenosyl methionine: structural inferences from variations on a theme and the reproducibility of SELEX.Nucleic Acids Res. 1997 May 15;25(10):2020-4. doi: 10.1093/nar/25.10.2020. Nucleic Acids Res. 1997. PMID: 9115371 Free PMC article.
-
A novel acidophilic RNA motif that recognizes coenzyme A.Biochemistry. 1998 Mar 31;37(13):4653-63. doi: 10.1021/bi972877p. Biochemistry. 1998. PMID: 9521786
-
Ribonucleopeptides: functional RNA-peptide complexes.Biopolymers. 2004;76(1):66-8. doi: 10.1002/bip.10576. Biopolymers. 2004. PMID: 14997476 Review.
-
In vitro selection methodologies to probe RNA function and structure.Mol Divers. 1995 Sep;1(1):69-78. doi: 10.1007/BF01715810. Mol Divers. 1995. PMID: 9237195 Review.
Cited by
-
A novel dengue virus detection method that couples DNAzyme and gold nanoparticle approaches.Virol J. 2013 Jun 28;10:201. doi: 10.1186/1743-422X-10-201. Virol J. 2013. PMID: 23809208 Free PMC article.
-
IPANEMAP: integrative probing analysis of nucleic acids empowered by multiple accessibility profiles.Nucleic Acids Res. 2020 Sep 4;48(15):8276-8289. doi: 10.1093/nar/gkaa607. Nucleic Acids Res. 2020. PMID: 32735675 Free PMC article.
-
Polypyrimidine-Tract-Binding Protein Isoforms Differentially Regulate the Hepatitis C Virus Internal Ribosome Entry Site.Viruses. 2022 Dec 20;15(1):8. doi: 10.3390/v15010008. Viruses. 2022. PMID: 36680049 Free PMC article.
-
RNA secondary structure of the feline immunodeficiency virus 5'UTR and Gag coding region.Nucleic Acids Res. 2008 Aug;36(14):4653-66. doi: 10.1093/nar/gkn447. Epub 2008 Jul 14. Nucleic Acids Res. 2008. PMID: 18625613 Free PMC article.
-
Mammalian RNA switches: Molecular rheostats in gene regulation, disease, and medicine.Comput Struct Biotechnol J. 2019 Oct 24;17:1326-1338. doi: 10.1016/j.csbj.2019.10.001. eCollection 2019. Comput Struct Biotechnol J. 2019. PMID: 31741723 Free PMC article. Review.
References
-
- Famulok M. (1999) Oligonucleotide aptamers that recognize small molecules. Curr. Opin. Struct. Biol., 9, 324–329. - PubMed
-
- Gold L., Polisky,B., Uhlenbeck,O. and Yarus,M. (1995) Diversity of oligonucleotide functions. Annu. Rev. Biochem., 64, 763–797. - PubMed
-
- Jayasena S.D. (1999) Aptamers: an emerging class of molecules that rival antibodies in diagnostics. Clin. Chem., 45, 1628–1650. - PubMed
-
- Brody E.N. and Gold,L. (2000) Aptamers as therapeutic and diagnostic agents. J. Biotechnol., 74, 5–13. - PubMed
-
- Burgstaller P., Jenne,A. and Blind,M. (2002) Aptamers and aptazymes: accelerating small molecule drug discovery. Curr. Opin. Drug Discov. Devel., 5, 690–700. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

