Characterization of the GRK2 binding site of Galphaq
- PMID: 15471870
- PMCID: PMC1432089
- DOI: 10.1074/jbc.M401438200
Characterization of the GRK2 binding site of Galphaq
Abstract
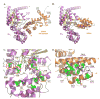
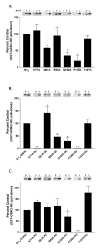
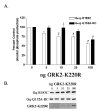
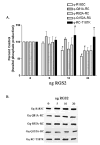
Heterotrimeric guanine nucleotide-binding proteins (G proteins) transmit signals from membrane bound G protein-coupled receptors (GPCRs) to intracellular effector proteins. The G(q) subfamily of Galpha subunits couples GPCR activation to the enzymatic activity of phospholipase C-beta (PLC-beta). Regulators of G protein signaling (RGS) proteins bind to activated Galpha subunits, including Galpha(q), and regulate Galpha signaling by acting as GTPase activating proteins (GAPs), increasing the rate of the intrinsic GTPase activity, or by acting as effector antagonists for Galpha subunits. GPCR kinases (GRKs) phosphorylate agonist-bound receptors in the first step of receptor desensitization. The amino termini of all GRKs contain an RGS homology (RH) domain, and binding of the GRK2 RH domain to Galpha(q) attenuates PLC-beta activity. The RH domain of GRK2 interacts with Galpha(q/11) through a novel Galpha binding surface termed the "C" site. Here, molecular modeling of the Galpha(q).GRK2 complex and site-directed mutagenesis of Galpha(q) were used to identify residues in Galpha(q) that interact with GRK2. The model identifies Pro(185) in Switch I of Galpha(q) as being at the crux of the interface, and mutation of this residue to lysine disrupts Galpha(q) binding to the GRK2-RH domain. Switch III also appears to play a role in GRK2 binding because the mutations Galpha(q)-V240A, Galpha(q)-D243A, both residues within Switch III, and Galpha(q)-Q152A, a residue that structurally supports Switch III, are defective in binding GRK2. Furthermore, GRK2-mediated inhibition of Galpha(q)-Q152A-R183C-stimulated inositol phosphate release is reduced in comparison to Galpha(q)-R183C. Interestingly, the model also predicts that residues in the helical domain of Galpha(q) interact with GRK2. In fact, the mutants Galpha(q)-K77A, Galpha(q)-L78D, Galpha(q)-Q81A, and Galpha(q)-R92A have reduced binding to the GRK2-RH domain. Finally, although the mutant Galpha(q)-T187K has greatly reduced binding to RGS2 and RGS4, it has little to no effect on binding to GRK2. Thus the RH domain A and C sites for Galpha(q) interaction rely on contacts with distinct regions and different Switch I residues in Galpha(q).
Figures


Similar articles
-
G protein-coupled receptor Kinase 2/G alpha q/11 interaction. A novel surface on a regulator of G protein signaling homology domain for binding G alpha subunits.J Biol Chem. 2003 Feb 21;278(8):6050-8. doi: 10.1074/jbc.M208787200. Epub 2002 Nov 8. J Biol Chem. 2003. PMID: 12427730
-
Selective regulation of Galpha(q/11) by an RGS domain in the G protein-coupled receptor kinase, GRK2.J Biol Chem. 1999 Nov 26;274(48):34483-92. doi: 10.1074/jbc.274.48.34483. J Biol Chem. 1999. PMID: 10567430
-
Inhibition of Galphaq-dependent PLC-beta1 activity by PKG and PKA is mediated by phosphorylation of RGS4 and GRK2.Am J Physiol Cell Physiol. 2007 Jan;292(1):C200-8. doi: 10.1152/ajpcell.00103.2006. Epub 2006 Aug 2. Am J Physiol Cell Physiol. 2007. PMID: 16885398
-
Application of RGS box proteins to evaluate G-protein selectivity in receptor-promoted signaling.Methods Enzymol. 2004;389:71-88. doi: 10.1016/S0076-6879(04)89005-0. Methods Enzymol. 2004. PMID: 15313560 Review.
-
Decoding Gαq signaling.Life Sci. 2016 May 1;152:99-106. doi: 10.1016/j.lfs.2016.03.037. Epub 2016 Mar 22. Life Sci. 2016. PMID: 27012764 Review.
Cited by
-
Adaptive evolution of signaling partners.Mol Biol Evol. 2015 Apr;32(4):998-1007. doi: 10.1093/molbev/msu404. Epub 2015 Jan 6. Mol Biol Evol. 2015. PMID: 25568345 Free PMC article.
-
The fifth element in animal Galpha protein evolution.Commun Integr Biol. 2009 May;2(3):227-9. doi: 10.4161/cib.2.3.8080. Commun Integr Biol. 2009. PMID: 19641738 Free PMC article.
-
Revealing the Activity of Trimeric G-proteins in Live Cells with a Versatile Biosensor Design.Cell. 2020 Aug 6;182(3):770-785.e16. doi: 10.1016/j.cell.2020.06.020. Epub 2020 Jul 6. Cell. 2020. PMID: 32634377 Free PMC article.
-
A peptide of the RGS domain of GRK2 binds and inhibits Gα(q) to suppress pathological cardiac hypertrophy and dysfunction.Sci Signal. 2016 Mar 22;9(420):ra30. doi: 10.1126/scisignal.aae0549. Sci Signal. 2016. PMID: 27016525 Free PMC article.
-
Noncanonical Roles of G Protein-coupled Receptor Kinases in Cardiovascular Signaling.J Cardiovasc Pharmacol. 2017 Sep;70(3):129-141. doi: 10.1097/FJC.0000000000000483. J Cardiovasc Pharmacol. 2017. PMID: 28328744 Free PMC article. Review.
References
-
- Bourne HR. Curr Opin Cell Biol. 1997;9:134–142. - PubMed
-
- Krupnick JG, Benovic JL. Annu Rev Pharmacol Toxicol. 1998;38:289–319. - PubMed
-
- Hausdorff WP, Caron MG, Lefkowitz RJ. FASEB J. 1990;4:2881–2889. - PubMed
-
- Goodman OB, Jr, Krupnick JG, Santini F, Gurevich VV, Penn RB, Gagnon AW, Keen JH, Benovic JL. Nature. 1996;383:447–450. - PubMed
-
- Ferguson SS, Downey WE, 3rd, Colapietro AM, Barak LS, Menard L, Caron MG. Science. 1996;271:363–366. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

