Modeling P-loops domain of sodium channel: homology with potassium channels and interaction with ligands
- PMID: 15475578
- PMCID: PMC1304997
- DOI: 10.1529/biophysj.104.048173
Modeling P-loops domain of sodium channel: homology with potassium channels and interaction with ligands
Abstract
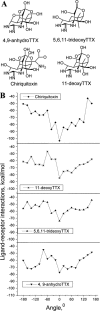
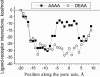
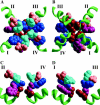
A large body of experimental data on Na+ channels is available, but the interpretation of these data in structural terms is difficult in the absence of a high-resolution structure. Essentially different electrophysiological and pharmacological properties of Na+ and K+ channels and poor identity of their sequences obstruct homology modeling of Na+ channels. In this work, we built the P-loops model of the Na+ channel, in which the pore helices are arranged exactly as in the MthK bacterial K+ channel. The conformation of the selectivity-filter region, which includes residues in positions -2 through +4 from the DEKA locus, was shaped around rigid molecules of saxitoxin and tetrodotoxin that are known to form multiple contacts with this region. Intensive Monte Carlo minimization that started from the MthK-like conformation produced practically identical saxitoxin- and tetrodotoxin-based models. The latter was tested to explain a wide range of experimental data that were not used at the model building stage. The docking of tetrodotoxin analogs unambiguously predicted their optimal orientation and the interaction energy that correlates with the experimental activity. The docking of mu-conotoxin produced a binding model consistent with experimentally known toxin-channel contacts. Monte Carlo-minimized energy profiles of tetramethylammonium pulled through the selectivity-filter region explain the paradoxical experimental data that this organic cation permeates via the DEAA but not the AAAA mutant of the DEKA locus. The model is also consistent with earlier proposed concepts on the Na+ channel selectivity as well as Ca2+ selectivity of the EEEE mutant of the DEKA locus. Thus, the model integrates available experimental data on the Na+ channel P-loops domain, and suggests that it is more similar to K+ channels than was believed before.
Figures



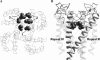
References
-
- Aqvist, J., and V. Luzhkov. 2000. Ion permeation mechanism of the potassium channel. Nature. 404:881–884. - PubMed
-
- Backx, P. H., D. T. Yue, J. H. Lawrence, E. Marban, and G. F. Tomaselli. 1992. Molecular localization of an ion-binding site within the pore of mammalian sodium channels. Science. 257:248–251. - PubMed
-
- Becker, S., E. Prusak-Sochaczewski, G. Zamponi, A. G. Beck-Sickinger, R. D. Gordon, and R. J. French. 1992. Action of derivatives of μ-conotoxin GIIIA on sodium channels. Single amino acid substitutions in the toxin separately affect association and dissociation rates. Biochemistry. 31:8229–8238. - PubMed
-
- Berneche, S., and B. Roux. 2001. Energetics of ion conduction through the K+ channel. Nature. 414:73–77. - PubMed
-
- Brooks, C. L., B. M. Pettitt, and M. Karplus. 1985. Structural and energetic effects of truncating long-ranged interactions in ionic polar fluids. J. Chem. Phys. 83:5897–5908.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials
Miscellaneous

