Regulation of PutA-membrane associations by flavin adenine dinucleotide reduction
- PMID: 15476410
- PMCID: PMC1513155
- DOI: 10.1021/bi048596g
Regulation of PutA-membrane associations by flavin adenine dinucleotide reduction
Abstract
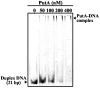
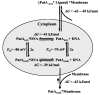
Proline utilization A (PutA) from Escherichia coli is a multifunctional flavoprotein that is both a transcriptional repressor of the proline utilization (put) genes and a membrane-associated enzyme which catalyzes the 4-electron oxidation of proline to glutamate. Previously, proline was shown to induce PutA-membrane binding and alter the intracellular location and function of PutA. To distinguish the roles of substrate binding and FAD reduction in the mechanism of how PutA changes from a DNA-binding protein to a membrane-bound enzyme, the kinetic parameters of PutA-membrane binding were measured under different conditions using model lipid bilayers and surface plasmon resonance (SPR). The effects of proline, FAD reduction, and proline analogues on PutA-membrane associations were determined. Oxidized PutA shows no binding to E. coli polar lipid vesicles. In contrast, proline and sodium dithionite induce tight binding of PutA to the lipid bilayer with indistinguishable kinetic parameters and an estimated dissociation constant (K(D)) of <0.01 nM (pH 7.4) for the reduced PutA-lipid complex. Proline analogues such as L-THFA and DL-P5C also stimulate PutA binding to E. coli polar lipid vesicles with K(D) values ranging from approximately 3.6 to 34 nM (pH 7.4) for the PutA-lipid complex. The greater PutA-membrane binding affinity (>300-fold) generated by FAD reduction relative to the nonreducing ligands demonstrates that FAD reduction controls PutA-membrane associations. On the basis of SPR kinetic analysis with differently charged lipid bilayers, the driving force for PutA-membrane binding is primarily hydrophobic. In the SPR experiments membrane-bound PutA did not bind put control DNA, confirming that the membrane-binding and DNA-binding activities of PutA are mutually exclusive. A model for the regulation of PutA is described in which the overall translocation of PutA from the cytoplasm to the membrane is driven by FAD reduction and the subsequent energy difference ( approximately 24 kJ/mol) between PutA-membrane and PutA-DNA binding.
Figures




Similar articles
-
Characterization of a bifunctional PutA homologue from Bradyrhizobium japonicum and identification of an active site residue that modulates proline reduction of the flavin adenine dinucleotide cofactor.Biochemistry. 2005 Jun 28;44(25):9130-9. doi: 10.1021/bi050629k. Biochemistry. 2005. PMID: 15966737 Free PMC article.
-
Flavin redox state triggers conformational changes in the PutA protein from Escherichia coli.Biochemistry. 2003 May 13;42(18):5469-77. doi: 10.1021/bi0272196. Biochemistry. 2003. PMID: 12731889
-
Redox properties of the PutA protein from Escherichia coli and the influence of the flavin redox state on PutA-DNA interactions.Biochemistry. 2001 Apr 17;40(15):4714-21. doi: 10.1021/bi0019491. Biochemistry. 2001. PMID: 11294639
-
Structure, function, and mechanism of proline utilization A (PutA).Arch Biochem Biophys. 2017 Oct 15;632:142-157. doi: 10.1016/j.abb.2017.07.005. Epub 2017 Jul 14. Arch Biochem Biophys. 2017. PMID: 28712849 Free PMC article. Review.
-
Direct linking of metabolism and gene expression in the proline utilization A protein from Escherichia coli.Amino Acids. 2008 Nov;35(4):711-8. doi: 10.1007/s00726-008-0053-6. Epub 2008 Mar 7. Amino Acids. 2008. PMID: 18324349 Free PMC article. Review.
Cited by
-
Role of Proline in Pathogen and Host Interactions.Antioxid Redox Signal. 2019 Feb 1;30(4):683-709. doi: 10.1089/ars.2017.7335. Epub 2018 Feb 2. Antioxid Redox Signal. 2019. PMID: 29241353 Free PMC article. Review.
-
Cloning, purification and crystallization of Thermus thermophilus proline dehydrogenase.Acta Crystallogr Sect F Struct Biol Cryst Commun. 2005 Aug 1;61(Pt 8):737-9. doi: 10.1107/S1744309105019779. Epub 2005 Jul 8. Acta Crystallogr Sect F Struct Biol Cryst Commun. 2005. PMID: 16511143 Free PMC article.
-
Insights into membrane association of Klebsiella pneumoniae NifL under nitrogen-fixing conditions from mutational analysis.J Bacteriol. 2011 Feb;193(3):695-705. doi: 10.1128/JB.00775-10. Epub 2010 Nov 5. J Bacteriol. 2011. PMID: 21057007 Free PMC article.
-
Steady-state kinetic mechanism of the proline:ubiquinone oxidoreductase activity of proline utilization A (PutA) from Escherichia coli.Arch Biochem Biophys. 2011 Dec 15;516(2):113-20. doi: 10.1016/j.abb.2011.10.011. Epub 2011 Oct 25. Arch Biochem Biophys. 2011. PMID: 22040654 Free PMC article.
-
Solution structure of the Pseudomonas putida protein PpPutA45 and its DNA complex.Proteins. 2009 Apr;75(1):12-27. doi: 10.1002/prot.22217. Proteins. 2009. PMID: 18767154 Free PMC article.
References
-
- Brown E, Wood JM. Redesigned purification yields a fully functional PutA protein dimer from Escherichia coli. J. Biol. Chem. 1992;267:13086–13092. - PubMed
-
- Abrahamson JLA, Baker LG, Stephenson JT, Wood JM. Proline dehydrogenase from Escherichia K12, properties of the membrane-associated enzyme. Eur. J. Biochem. 1983;134:77–82. - PubMed
-
- Graham S, Stephenson JT, Wood JM. Proline dehydrogenase from Escherichia coli K12, reconstitution of functional membrane association. J. Biol. Chem. 1984;259:2656–2661. - PubMed
-
- Ling M, Allen SW, Wood JM. Sequence analysis identifies the proline dehydrogenase and pyrroline-5-carboxylate dehydrogenase domains of the multifunctional Escherichia coli PutA protein. J. Mol. Biol. 1994;245:950–956. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

