Protein kinases of the human malaria parasite Plasmodium falciparum: the kinome of a divergent eukaryote
- PMID: 15479470
- PMCID: PMC526369
- DOI: 10.1186/1471-2164-5-79
Protein kinases of the human malaria parasite Plasmodium falciparum: the kinome of a divergent eukaryote
Abstract
Background: Malaria, caused by the parasitic protist Plasmodium falciparum, represents a major public health problem in the developing world. The P. falciparum genome has been sequenced, which provides new opportunities for the identification of novel drug targets. Eukaryotic protein kinases (ePKs) form a large family of enzymes with crucial roles in most cellular processes; hence malarial ePKS represent potential drug targets. We report an exhaustive analysis of the P. falciparum genomic database (PlasmoDB) aimed at identifying and classifying all ePKs in this organism.
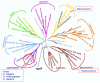
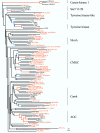
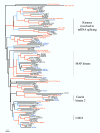
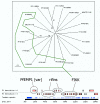
Results: Using a variety of bioinformatics tools, we identified 65 malarial ePK sequences and constructed a phylogenetic tree to position these sequences relative to the seven established ePK groups. Predominant features of the tree were: (i) that several malarial sequences did not cluster within any of the known ePK groups; (ii) that the CMGC group, whose members are usually involved in the control of cell proliferation, had the highest number of malarial ePKs; and (iii) that no malarial ePK clustered with the tyrosine kinase (TyrK) or STE groups, pointing to the absence of three-component MAPK modules in the parasite. A novel family of 20 ePK-related sequences was identified and called FIKK, on the basis of a conserved amino acid motif. The FIKK family seems restricted to Apicomplexa, with 20 members in P. falciparum and just one member in some other Apicomplexan species.
Conclusion: The considerable phylogenetic distance between Apicomplexa and other Eukaryotes is reflected by profound divergences between the kinome of malaria parasites and that of yeast or mammalian cells.
Figures


References
-
- Doerig C. Protein kinases as targets for anti-parasitic chemotherapy. Biochim Biophys Acta. 2004:155–168. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials

