The amino terminus of Epstein-Barr Virus (EBV) nuclear antigen 1 contains AT hooks that facilitate the replication and partitioning of latent EBV genomes by tethering them to cellular chromosomes
- PMID: 15479791
- PMCID: PMC523237
- DOI: 10.1128/JVI.78.21.11487-11505.2004
The amino terminus of Epstein-Barr Virus (EBV) nuclear antigen 1 contains AT hooks that facilitate the replication and partitioning of latent EBV genomes by tethering them to cellular chromosomes
Abstract
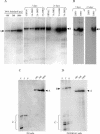
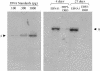
During latency, Epstein-Barr virus (EBV) is stably maintained as a circular plasmid that is replicated once per cell cycle and partitioned at mitosis. Both these processes require a single viral protein, EBV nuclear antigen 1 (EBNA1), which binds two clusters of cognate binding sites within the latent viral origin, oriP. EBNA1 is known to associate with cellular metaphase chromosomes through chromosome-binding domains within its amino terminus, an association that we have determined to be required not only for the partitioning of oriP plasmids but also for their replication. One of the chromosome-binding domains of EBNA1 associates with a cellular nucleolar protein, EBP2, and it has been proposed that this interaction underlies that ability of EBNA1 to bind metaphase chromosomes. Here we demonstrate that EBNA1's chromosome-binding domains are AT hooks, a DNA-binding motif found in a family of proteins that bind the scaffold-associated regions on metaphase chromosomes. Further, we demonstrate that the ability of EBNA1 to stably replicate and partition oriP plasmids correlates with its AT hook activity and not its association with EBP2. Finally, we examine the contributions of EBP2 toward the ability of EBNA1 to associate with metaphase chromosomes in human cells, as well as support the replication and partitioning of oriP plasmids in human cells. Our results indicate that it is unlikely that EBP2 directly mediates these activities of EBNA1 in human cells.
Figures
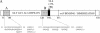
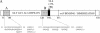




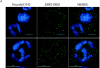



References
-
- Aiyar, A., and B. Sugden. 1998. Fusions between Epstein-Barr viral nuclear antigen-1 of Epstein-Barr virus and the large T-antigen of simian virus 40 replicate their cognate origins. J. Biol. Chem. 273:33073-33081. - PubMed
-
- Banks, G. C., B. Mohr, and R. Reeves. 1999. The HMG-I(Y) A.T-hook peptide motif confers DNA-binding specificity to a structured chimeric protein. J. Biol. Chem. 274:16536-16544. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources

