Role of Unc104/KIF1-related motor proteins in mitochondrial transport in Neurospora crassa
- PMID: 15483054
- PMCID: PMC539160
- DOI: 10.1091/mbc.e04-05-0413
Role of Unc104/KIF1-related motor proteins in mitochondrial transport in Neurospora crassa
Abstract
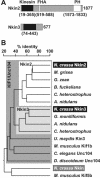
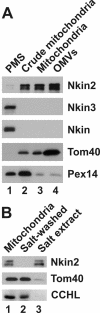
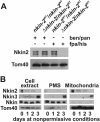
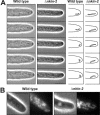
Eukaryotic cells use diverse cytoskeleton-dependent machineries to control inheritance and intracellular positioning of mitochondria. In particular, microtubules play a major role in mitochondrial motility in the filamentous fungus Neurospora crassa and in mammalian cells. We examined the role of two novel Unc104/KIF1-related members of the kinesin family, Nkin2 and Nkin3, in mitochondrial motility in Neurospora. The Nkin2 protein is required for mitochondrial interactions with microtubules in vitro. Mutant hyphae lacking Nkin2 show mitochondrial motility defects in vivo early after germination of conidiospores. Nkin3, a member of a unique fungal-specific subgroup of small Unc104/KIF1-related proteins, is not associated with mitochondria in wild-type cells. However, it is highly expressed and recruited to mitochondria in Deltankin-2 mutants. Mitochondria lacking Nkin2 require Nkin3 for binding to microtubules in vitro, and mitochondrial motility defects in Deltankin-2 mutants disappear with up-regulation of Nkin3 in vivo. We propose that mitochondrial transport is mediated by Nkin2 in Neurospora, and organelle motility defects in Deltankin-2 mutants are rescued by Nkin3. Apparently, a highly versatile complement of organelle motors allows the cell to efficiently respond to exogenous challenges, a process that might also account for the great variety of different mitochondrial transport systems that have evolved in eukaryotic cells.
Figures



Similar articles
-
Neurospora crassa NKIN2, a kinesin-3 motor, transports early endosomes and is required for polarized growth.Eukaryot Cell. 2013 Jul;12(7):1020-32. doi: 10.1128/EC.00081-13. Epub 2013 May 17. Eukaryot Cell. 2013. PMID: 23687116 Free PMC article.
-
Microtubules and associated molecular motors in Neurospora crassa.Mycologia. 2016 May-Jun;108(3):515-27. doi: 10.3852/15-323. Epub 2016 Mar 7. Mycologia. 2016. PMID: 26951369
-
Interaction of mitochondria with microtubules in the filamentous fungus Neurospora crassa.J Cell Sci. 2002 May 1;115(Pt 9):1931-7. doi: 10.1242/jcs.115.9.1931. J Cell Sci. 2002. PMID: 11956324
-
Mitochondrial dynamics in filamentous fungi.Fungal Genet Biol. 2002 Jul;36(2):91-7. doi: 10.1016/S1087-1845(02)00019-1. Fungal Genet Biol. 2002. PMID: 12081462 Review.
-
Structural and functional organization of growing tips of Neurospora crassa Hyphae.Biochemistry (Mosc). 2014 Jul;79(7):593-607. doi: 10.1134/S0006297914070025. Biochemistry (Mosc). 2014. PMID: 25108323 Review.
Cited by
-
The class V myosin motor protein, Myo2, plays a major role in mitochondrial motility in Saccharomyces cerevisiae.J Cell Biol. 2008 Apr 7;181(1):119-30. doi: 10.1083/jcb.200709099. J Cell Biol. 2008. PMID: 18391073 Free PMC article.
-
Hyphal growth: a tale of motors, lipids, and the Spitzenkörper.Eukaryot Cell. 2007 Mar;6(3):351-60. doi: 10.1128/EC.00381-06. Epub 2007 Jan 26. Eukaryot Cell. 2007. PMID: 17259546 Free PMC article. Review. No abstract available.
-
Intracellular Cargo Transport by Kinesin-3 Motors.Biochemistry (Mosc). 2017 Jul;82(7):803-815. doi: 10.1134/S0006297917070057. Biochemistry (Mosc). 2017. PMID: 28918744 Free PMC article. Review.
-
Mitochondrial DNA-depleted A549 cells are resistant to bleomycin.Am J Physiol Lung Cell Mol Physiol. 2012 Sep;303(5):L413-24. doi: 10.1152/ajplung.00343.2011. Epub 2012 Jul 6. Am J Physiol Lung Cell Mol Physiol. 2012. PMID: 22773697 Free PMC article.
-
CLASP regulates mitochondrial distribution in Schizosaccharomyces pombe.J Cell Biol. 2008 Jul 14;182(1):41-9. doi: 10.1083/jcb.200712147. Epub 2008 Jul 7. J Cell Biol. 2008. PMID: 18606849 Free PMC article.
References
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

