Notch subunit heterodimerization and prevention of ligand-independent proteolytic activation depend, respectively, on a novel domain and the LNR repeats
- PMID: 15485896
- PMCID: PMC522238
- DOI: 10.1128/MCB.24.21.9265-9273.2004
Notch subunit heterodimerization and prevention of ligand-independent proteolytic activation depend, respectively, on a novel domain and the LNR repeats
Abstract
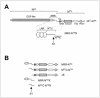
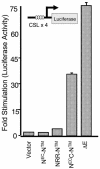
Notch proteins are transmembrane receptors that participate in a highly conserved signaling pathway that regulates morphogenesis in metazoans. Newly synthesized Notch receptors are proteolytically cleaved during transit to the cell surface, creating heterodimeric mature receptors comprising noncovalently associated extracellular (N(EC)) and transmembrane (N) subunits. Ligand binding activates Notch by inducing two successive proteolytic cleavages, catalyzed by metalloproteases and gamma-secretase, respectively, that permit the intracellular portion of N to translocate to the nucleus and activate transcription of target genes. Prior work has shown that the presence of N(EC) prevents ligand-independent activation of N, but the mechanisms involved are poorly understood. Here, we define the roles of two regions at the C-terminal end of N(EC) that participate in maintaining the integrity of resting Notch receptors through distinct mechanisms. The first region, a hydrophobic, previously uncharacterized portion of N(EC), is sufficient to form stable complexes with the extracellular portion of N. The second region, consisting of the three Lin12/Notch repeats, is not needed for heterodimerization but acts to protect N from ligand-independent cleavage by metalloproteases. Together, these two contiguous regions of N(EC) impose crucial restraints that prevent premature Notch receptor activation.
Figures





References
-
- Artavanis-Tsakonas, S., M. D. Rand, and R. J. Lake. 1999. Notch signaling: cell fate control and signal integration in development. Science 284:770-776. - PubMed
-
- Aster, J., W. Pear, R. Hasserjian, H. Erba, F. Davi, B. Luo, M. Scott, D. Baltimore, and J. Sklar. 1994. Functional analysis of the TAN-1 gene, a human homolog of Drosophila notch. Cold Spring Harbor Symp. Quant. Biol. 59:125-136. - PubMed
-
- Aster, J. C., W. B. Simms, Z. Zavala-Ruiz, V. Patriub, C. L. North, and S. C. Blacklow. 1999. The folding and structural integrity of the first LIN-12 module of human Notch1 are calcium-dependent. Biochemistry 38:4736-4742. - PubMed
-
- Bellavia, D., A. F. Campese, E. Alesse, A. Vacca, M. P. Felli, A. Balestri, A. Stoppacciaro, C. Tiveron, L. Tatangelo, M. Giovarelli, C. Gaetano, L. Ruco, E. S. Hoffman, A. C. Hayday, U. Lendahl, L. Frati, A. Gulino, and I. Screpanti. 2000. Constitutive activation of NF-kappaB and T-cell leukemia/lymphoma in Notch3 transgenic mice. EMBO J. 19:3337-3348. - PMC - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
