Interferon-inducible ubiquitin E2, Ubc8, is a conjugating enzyme for protein ISGylation
- PMID: 15485925
- PMCID: PMC522249
- DOI: 10.1128/MCB.24.21.9592-9600.2004
Interferon-inducible ubiquitin E2, Ubc8, is a conjugating enzyme for protein ISGylation
Abstract
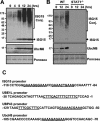
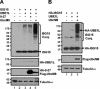
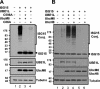
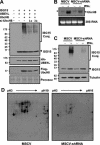
Protein ISGylation is unique among ubiquitin-like conjugation systems in that the expression and conjugation processes are induced by specific stimuli, mainly via the alpha/beta interferon signaling pathway. It has been suggested that protein ISGylation plays a special role in the immune response, because of its interferon-signal dependency and its appearance only in higher eukaryotic organisms. Here, we report the identification of an ISG15-conjugating enzyme, Ubc8. Like other components of the protein ISGylation system (ISG15, UBE1L, and UBP43), Ubc8 is an interferon-inducible protein. Ubc8 clearly mediates protein ISGylation in transfection assays. The reduction of Ubc8 expression by small interfering RNA causes a decrease in protein ISGylation in HeLa cells upon interferon treatment. Neither UbcH7/UbcM4, the closest homologue of Ubc8 among known ubiquitin E2s, nor the small ubiquitin-like modifier E2 Ubc9 supports protein ISGylation. These findings strongly suggest that Ubc8 is a major ISG15-conjugating enzyme responsible for protein ISGylation upon interferon stimulation. Furthermore, we established an assay system to detect ISGylated target proteins by cotransfection of ISG15, UBE1L, and Ubc8 together with a target protein to be analyzed. This method provides an easy and effective way to identify new targets for the ISGylation system and will facilitate related studies.
Figures



References
-
- Chen, C. D., D. S. Welsbie, C. Tran, S. H. Baek, R. Chen, R. Vessella, M. G. Rosenfeld, and C. L. Sawyers. 2004. Molecular determinants of resistance to antiandrogen therapy. Nat. Med. 10:33-39. - PubMed
-
- Chin, L. S., J. P. Vavalle, and L. Li. 2002. Staring, a novel E3 ubiquitin-protein ligase that targets syntaxin 1 for degradation. J. Biol. Chem. 277:35071-35079. - PubMed
-
- Chung, C. H., and S. H. Baek. 1999. Deubiquitinating enzymes: their diversity and emerging roles. Biochem. Biophys. Res. Commun. 266:633-640. - PubMed
-
- D'Andrea, A., and D. Pellman. 1998. Deubiquitinating enzymes: a new class of biological regulators. Crit. Rev. Biochem. Mol. Biol. 33:337-352. - PubMed
-
- Darnell, J. E., Jr., I. M. Kerr, and G. R. Stark. 1994. Jak-STAT pathways and transcriptional activation in response to IFNs and other extracellular signaling proteins. Science 264:1415-1421. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous
