The sequences and activities of RegB endoribonucleases of T4-related bacteriophages
- PMID: 15486207
- PMCID: PMC524301
- DOI: 10.1093/nar/gkh892
The sequences and activities of RegB endoribonucleases of T4-related bacteriophages
Abstract
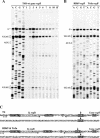
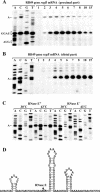
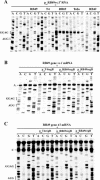
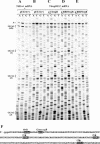
The RegB endoribonuclease encoded by bacteriophage T4 is a unique sequence-specific nuclease that cleaves in the middle of GGAG or, in a few cases, GGAU tetranucleotides, preferentially those found in the Shine-Dalgarno regions of early phage mRNAs. In this study, we examined the primary structures and functional properties of RegB ribonucleases encoded by T4-related bacteriophages. We show that all but one of 36 phages tested harbor the regB gene homologues and the similar signals for transcriptional and post-transcriptional autogenous regulation of regB expression. Phage RB49 in addition to gpRegB utilizes Escherichia coli endoribonuclease E for the degradation of its transcripts for gene regB. The deduced primary structure of RegB proteins of 32 phages studied is almost identical to that of T4, while the sequences of RegB encoded by phages RB69, TuIa and RB49 show substantial divergence from their T4 counterpart. Functional studies using plasmid-phage systems indicate that RegB nucleases of phages T4, RB69, TuIa and RB49 exhibit different activity towards GGAG and GGAU motifs in the specific locations. We expect that the availability of the different phylogenetic variants of RegB may help to localize the amino acid determinants that contribute to the specificity and cleavage efficiency of this processing enzyme.
Figures



Similar articles
-
Endoribonuclease RegB from bacteriophage T4 is necessary for the degradation of early but not middle or late mRNAs.J Mol Biol. 2000 Apr 14;297(5):1063-74. doi: 10.1006/jmbi.2000.3626. J Mol Biol. 2000. PMID: 10764573
-
Dual role of the sequence-specific bacteriophage T4 endoribonuclease RegB. mRNA inactivation and mRNA destabilization.J Mol Biol. 1993 Oct 5;233(3):429-46. doi: 10.1006/jmbi.1993.1522. J Mol Biol. 1993. PMID: 8411154
-
Transcription and RNA processing during expression of genes preceding DNA ligase gene 30 in T4-related bacteriophages.Virology. 2006 Jan 20;344(2):378-90. doi: 10.1016/j.virol.2005.09.001. Epub 2005 Oct 12. Virology. 2006. PMID: 16225899
-
Post-transcriptional controls in bacteriophage T4: roles of the sequence-specific endoribonuclease RegB.FEMS Microbiol Rev. 1995 Aug;17(1-2):141-50. doi: 10.1111/j.1574-6976.1995.tb00196.x. FEMS Microbiol Rev. 1995. PMID: 7669340 Review.
-
RNA processing and decay in bacteriophage T4.Prog Mol Biol Transl Sci. 2009;85:43-89. doi: 10.1016/S0079-6603(08)00802-7. Prog Mol Biol Transl Sci. 2009. PMID: 19215770 Review.
Cited by
-
Activation of RegB endoribonuclease by S1 ribosomal protein requires an 11 nt conserved sequence.Nucleic Acids Res. 2006;34(22):6549-60. doi: 10.1093/nar/gkl911. Epub 2006 Nov 27. Nucleic Acids Res. 2006. PMID: 17130171 Free PMC article.
-
Post-transcriptional control by bacteriophage T4: mRNA decay and inhibition of translation initiation.Virol J. 2010 Dec 3;7:360. doi: 10.1186/1743-422X-7-360. Virol J. 2010. PMID: 21129205 Free PMC article. Review.
-
Shine-Dalgarno sequence of bacteriophage T4: GAGG prevails in early genes.Mol Biol Rep. 2012 Jan;39(1):33-9. doi: 10.1007/s11033-011-0707-4. Epub 2011 Apr 30. Mol Biol Rep. 2012. PMID: 21533668
References
-
- Deutscher M.P. and Li,Z. (2001) Exoribonucleases and their multiple roles in RNA metabolism. Prog. Nucleic Acid Res. Mol. Biol., 66, 67–105. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Other Literature Sources

