ACCUMULATION OF PHOTOSYSTEM ONE1, a member of a novel gene family, is required for accumulation of [4Fe-4S] cluster-containing chloroplast complexes and antenna proteins
- PMID: 15494558
- PMCID: PMC527200
- DOI: 10.1105/tpc.104.024935
ACCUMULATION OF PHOTOSYSTEM ONE1, a member of a novel gene family, is required for accumulation of [4Fe-4S] cluster-containing chloroplast complexes and antenna proteins
Abstract
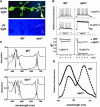
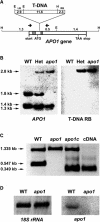
To investigate the nuclear-controlled mechanisms of [4Fe-4S] cluster assembly in chloroplasts, we selected Arabidopsis thaliana mutants with a decreased content of photosystem I (PSI) containing three [4Fe-4S] clusters. One identified gene, ACCUMULATION OF PHOTOSYSTEM ONE1 (APO1), belongs to a previously unknown gene family with four defined groups (APO1 to APO4) only found in nuclear genomes of vascular plants. All homologs contain two related motifs of approximately 100 amino acid residues that could potentially provide ligands for [4Fe-4S] clusters. APO1 is essentially required for photoautotrophic growth, and levels of PSI core subunits are below the limit of detection in the apo1 mutant. Unlike other Arabidopsis PSI mutants, apo1 fails to accumulate significant amounts of the outer antenna subunits of PSI and PSII and to form grana stacks. In particular, APO1 is essentially required for stable accumulation of other plastid-encoded and nuclear-encoded [4Fe-4S] cluster complexes within the chloroplast, whereas [2Fe-2S] cluster-containing complexes appear to be unaffected. In vivo labeling experiments and analyses of polysome association suggest that translational elongation of the PSI transcripts psaA and psaB is specifically arrested in the mutant. Taken together, our findings suggest that APO1 is involved in the stable assembly of several [4Fe-4S] cluster-containing complexes of chloroplasts and interferes with translational events probably in association with plastid nucleoids.
Figures





References
-
- Barkan, A. (1998). Approaches to investigating nuclear genes that function in chloroplast biogenesis in land plants. Methods Enzymol. 297, 38–57.
-
- Barkan, A., and Goldschmidt-Clermont, M. (2000). Participation of nuclear genes in chloroplast gene expression. Biochimie 82, 559–572. - PubMed
-
- Beinert, H., Holm, R.H., and Munck, E. (2000). Iron-sulfur clusters: Nature's modular, multipurpose structures. Science 277, 653–659. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

