Differential expression of porins OmpP2A and OmpP2B of Haemophilus ducreyi
- PMID: 15501753
- PMCID: PMC523061
- DOI: 10.1128/IAI.72.11.6271-6278.2004
Differential expression of porins OmpP2A and OmpP2B of Haemophilus ducreyi
Abstract
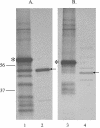
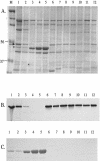
Haemophilus ducreyi is a strict human pathogen and the causative agent of the sexually transmitted disease chancroid. The genome of the human-passaged strain of H. ducreyi (35000HP) contains two homologous genes whose protein products have estimated molecular masses of 46 and 43 kDa. A comparative analysis of the deduced amino acid sequences revealed that these proteins share 27 to 33% identity to the outer membrane protein P2 (OmpP2), a major porin of Haemophilus influenzae. Therefore, these proteins have been designated OmpP2A and OmpP2B, respectively. The detection of ompP2A and ompP2B transcripts by reverse transcriptase PCR indicated that these genes were independently transcribed in H. ducreyi 35000HP. Western blot analysis of outer membrane proteins isolated from a geographically diverse collection of H. ducreyi clinical isolates revealed that OmpP2A and OmpP2B were differentially expressed among these strains. Although PCR analysis suggested that ompP2A and ompP2B were conserved among the strains tested, the differential expression observed was due to nucleotide additions and partial gene deletions. Purified OmpP2A and OmpP2B were isolated under nondenaturing conditions, and subsequent analysis demonstrated that these two proteins exhibited porin activity. OmpP2A and OmpP2B are the first porins described for H. ducreyi.
Figures




Similar articles
-
Comparative proteomic analysis of the Haemophilus ducreyi porin-deficient mutant 35000HP::P2AB.J Bacteriol. 2009 Apr;191(7):2144-52. doi: 10.1128/JB.01487-08. Epub 2008 Dec 19. J Bacteriol. 2009. PMID: 19103932 Free PMC article.
-
Expression of OmpP2A and OmpP2B is not required for pustule formation by Haemophilus ducreyi in human volunteers.Microb Pathog. 2006 Mar;40(3):110-5. doi: 10.1016/j.micpath.2005.11.005. Epub 2006 Jan 19. Microb Pathog. 2006. PMID: 16426809
-
The LspB protein is involved in the secretion of the LspA1 and LspA2 proteins by Haemophilus ducreyi.Infect Immun. 2004 Apr;72(4):1874-84. doi: 10.1128/IAI.72.4.1874-1884.2004. Infect Immun. 2004. PMID: 15039306 Free PMC article.
-
Biology of Haemophilus ducreyi.Microbiol Rev. 1989 Dec;53(4):377-89. doi: 10.1128/mr.53.4.377-389.1989. Microbiol Rev. 1989. PMID: 2687678 Free PMC article. Review.
-
Structure and function of the porin channel.Kidney Int. 1995 Oct;48(4):930-40. doi: 10.1038/ki.1995.374. Kidney Int. 1995. PMID: 8569102 Review. No abstract available.
Cited by
-
Activation of the CpxRA system by deletion of cpxA impairs the ability of Haemophilus ducreyi to infect humans.Infect Immun. 2010 Sep;78(9):3898-904. doi: 10.1128/IAI.00432-10. Epub 2010 Jul 6. Infect Immun. 2010. PMID: 20605985 Free PMC article.
-
Comparative proteomic analysis of the Haemophilus ducreyi porin-deficient mutant 35000HP::P2AB.J Bacteriol. 2009 Apr;191(7):2144-52. doi: 10.1128/JB.01487-08. Epub 2008 Dec 19. J Bacteriol. 2009. PMID: 19103932 Free PMC article.
-
A (p)ppGpp-null mutant of Haemophilus ducreyi is partially attenuated in humans due to multiple conflicting phenotypes.Infect Immun. 2014 Aug;82(8):3492-502. doi: 10.1128/IAI.01994-14. Epub 2014 Jun 9. Infect Immun. 2014. PMID: 24914217 Free PMC article.
-
Regulation of expression of the Haemophilus ducreyi LspB and LspA2 proteins by CpxR.Infect Immun. 2009 Aug;77(8):3402-11. doi: 10.1128/IAI.00292-09. Epub 2009 May 18. Infect Immun. 2009. PMID: 19451237 Free PMC article.
-
Characterization and comparative analysis of the genes encoding Haemophilus parasuis outer membrane proteins P2 and P5.J Bacteriol. 2009 Oct;191(19):5988-6002. doi: 10.1128/JB.00469-09. Epub 2009 Jul 24. J Bacteriol. 2009. PMID: 19633080 Free PMC article.
References
-
- Achouak, W., T. Heulin, and J. M. Pages. 2001. Multiple facets of bacterial porins. FEMS Microbiol. Lett. 199:1-7. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

