Quantitative imaging of lymphocyte membrane protein reorganization and signaling
- PMID: 15501943
- PMCID: PMC1305035
- DOI: 10.1529/biophysj.104.048827
Quantitative imaging of lymphocyte membrane protein reorganization and signaling
Abstract
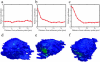
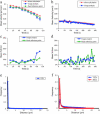
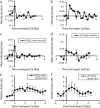
Changes in membrane protein localization are critical to establishing cell polarity and regulating cell signaling. Fluorescence microscopy of labeled proteins allows visualization of these changes, but quantitative analysis is needed to study this aspect of cell signaling in full mechanistic detail. We have developed a novel approach for quantitative assessment of membrane protein redistribution based on four-dimensional video microscopy of fluorescently labeled proteins. Our analytic system provides robust automated methods for cell surface reconstruction, cell shape tracking, cell-surface distance measurement, and cluster formation analysis. These methods permit statistical analyses and testing of mechanistic hypotheses regarding cell signaling. We have used this approach to measure antigen-dependent clustering of signaling molecules in CD4+ T lymphocytes, obtaining clustering velocities consistent with single-particle tracking data. Our system captures quantitative differences in clustering between signaling proteins with distinct biological functions. Our methods can be generalized to a range of cell-signaling phenomena and enable novel applications not feasible with single-particle studies, such as analysis of subcellular protein localization in live organ culture.
Figures



Similar articles
-
Optical techniques for imaging membrane domains in live cells (live-cell palm of protein clustering).Methods Enzymol. 2012;504:221-35. doi: 10.1016/B978-0-12-391857-4.00011-2. Methods Enzymol. 2012. PMID: 22264537 Review.
-
Quantifying protein densities on cell membranes using super-resolution optical fluctuation imaging.Nat Commun. 2017 Nov 23;8(1):1731. doi: 10.1038/s41467-017-01857-x. Nat Commun. 2017. PMID: 29170394 Free PMC article.
-
Roles of p-ERM and Rho-ROCK signaling in lymphocyte polarity and uropod formation.J Cell Biol. 2004 Oct 25;167(2):327-37. doi: 10.1083/jcb.200403091. J Cell Biol. 2004. PMID: 15504914 Free PMC article.
-
High-resolution multicolor imaging of dynamic signaling complexes in T cells stimulated by planar substrates.Sci STKE. 2003 Apr 8;2003(177):PL8. doi: 10.1126/stke.2003.177.pl8. Sci STKE. 2003. PMID: 12684528
-
Quantification and its applications in fluorescent microscopy imaging.Traffic. 2009 Aug;10(8):951-61. doi: 10.1111/j.1600-0854.2009.00938.x. Epub 2009 May 5. Traffic. 2009. PMID: 19500318 Review.
Cited by
-
ON IDENTIFYING INFORMATION FROM IMAGE-BASED SPATIAL POLARITY PHENOTYPES IN NEUTROPHILS.Proc IEEE Int Symp Biomed Imaging. 2010 Apr 1;14-17:1029-1032. doi: 10.1109/ISBI.2010.5490165. Proc IEEE Int Symp Biomed Imaging. 2010. PMID: 20725643 Free PMC article.
-
Synapse-directed delivery of immunomodulators using T-cell-conjugated nanoparticles.Biomaterials. 2012 Aug;33(23):5776-87. doi: 10.1016/j.biomaterials.2012.04.029. Epub 2012 May 15. Biomaterials. 2012. PMID: 22594972 Free PMC article.
References
-
- Agard, D. A., Y. Hiraoka, P. Shaw, and J. W. Sedat. 1989. Fluorescence microscopy in three dimensions. Methods Cell Biol. 30:353–377. - PubMed
-
- Boldin, M. P., I. L. Mett, E. E. Varfolomeev, I. Chumakov, Y. Shemer-Avni, J. H. Camonis, and D. Wallach. 1995. Self-association of the “death domains” of the p55 tumor necrosis factor (TNF) receptor and Fas/APO1 prompts signaling for TNF and Fas/APO1 effects. J. Biol. Chem. 270:387–391. - PubMed
-
- Bousso, P., N. R. Bhakta, R. S. Lewis, and E. Robey. 2002. Dynamics of thymocyte-stromal cell interactions visualized by two-photon microscopy. Science. 296:1876–1880. - PubMed
-
- Brillinger, D. R. 1997. A particle migrating randomly on a sphere. J. Theor. Probab. 10:429–443.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials

