A stromal Hsp100 protein is required for normal chloroplast development and function in Arabidopsis
- PMID: 15516497
- PMCID: PMC527159
- DOI: 10.1104/pp.104.052928
A stromal Hsp100 protein is required for normal chloroplast development and function in Arabidopsis
Abstract
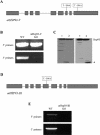
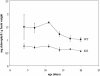
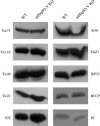
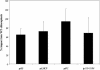
Molecular chaperones are required for the translocation of many proteins across organellar membranes, presumably by providing energy in the form of ATP hydrolysis for protein movement. In the chloroplast protein import system, a heat shock protein 100 (Hsp100), known as Hsp93, is hypothesized to be the chaperone providing energy for precursor translocation, although there is little direct evidence for this hypothesis. To learn more about the possible function of Hsp93 during protein import into chloroplasts, we isolated knockout mutant lines that contain T-DNA disruptions in either atHSP93-V or atHSP93-III, which encode the two Arabidopsis (Arabidopsis thaliana) homologs of Hsp93. atHsp93-V mutant plants are much smaller and paler than wild-type plants. In addition, mutant chloroplasts contain less thylakoid membrane when compared to the wild type. Plastid protein composition, however, seems to be largely unaffected in atHsp93-V knockout plants. Chloroplasts isolated from the atHsp93-V knockout mutant line are still able to import a variety of precursor proteins, but the rate of import of some of these precursors is significantly reduced. These results indicate that atHsp93-V has an important, but not essential, role in the biogenesis of Arabidopsis chloroplasts. In contrast, knockout mutant plants for atHsp93-III, the second Arabidopsis Hsp93 homolog, had a visible phenotype identical to the wild type, suggesting that atHsp93-III may not play as important a role as atHsp93-V in chloroplast development and/or function.
Figures


Similar articles
-
Further in vivo studies on the role of the molecular chaperone, Hsp93, in plastid protein import.Plant J. 2007 Apr;50(2):364-79. doi: 10.1111/j.1365-313X.2007.03060.x. Epub 2007 Mar 21. Plant J. 2007. PMID: 17376159
-
In vivo studies on the roles of Tic110, Tic40 and Hsp93 during chloroplast protein import.Plant J. 2005 Feb;41(3):412-28. doi: 10.1111/j.1365-313X.2004.02307.x. Plant J. 2005. PMID: 15659100
-
Functional Analysis of the Hsp93/ClpC Chaperone at the Chloroplast Envelope.Plant Physiol. 2016 Jan;170(1):147-62. doi: 10.1104/pp.15.01538. Epub 2015 Nov 19. Plant Physiol. 2016. PMID: 26586836 Free PMC article.
-
Molecular chaperone involvement in chloroplast protein import.Biochim Biophys Acta. 2013 Feb;1833(2):332-40. doi: 10.1016/j.bbamcr.2012.03.019. Epub 2012 Apr 12. Biochim Biophys Acta. 2013. PMID: 22521451 Review.
-
Molecular chaperones involved in chloroplast protein import.Biochim Biophys Acta. 2001 Dec 12;1541(1-2):102-13. doi: 10.1016/s0167-4889(01)00148-3. Biochim Biophys Acta. 2001. PMID: 11750666 Review.
Cited by
-
The motors of protein import into chloroplasts.Plant Signal Behav. 2011 Sep;6(9):1397-401. doi: 10.4161/psb.6.9.16916. Plant Signal Behav. 2011. PMID: 22019640 Free PMC article.
-
Quantitative analysis of the chloroplast molecular chaperone ClpC/Hsp93 in Arabidopsis reveals new insights into its localization, interaction with the Clp proteolytic core, and functional importance.J Biol Chem. 2014 Apr 18;289(16):11318-11330. doi: 10.1074/jbc.M113.534552. Epub 2014 Mar 5. J Biol Chem. 2014. PMID: 24599948 Free PMC article.
-
A Protochlorophyllide (Pchlide) a Oxygenase for Plant Viability.Front Plant Sci. 2019 May 15;10:593. doi: 10.3389/fpls.2019.00593. eCollection 2019. Front Plant Sci. 2019. PMID: 31156665 Free PMC article.
-
Accumulation of high contents of free amino acids in the leaves of Nicotiana benthamiana by the co-suppression of NbClpC1 and NbClpC2 genes.Plant Cell Rep. 2015 Mar;34(3):355-65. doi: 10.1007/s00299-014-1714-4. Epub 2014 Nov 30. Plant Cell Rep. 2015. PMID: 25433858
-
The TOC GTPase Receptors: Regulators of the Fidelity, Specificity and Substrate Profiles of the General Protein Import Machinery of Chloroplasts.Protein J. 2019 Jun;38(3):343-350. doi: 10.1007/s10930-019-09846-3. Protein J. 2019. PMID: 31201619 Free PMC article. Review.
References
-
- Bauerle C, Keegstra K (1991) Full-length plastocyanin precursor is translocated across isolated thylakoid membranes. J Biol Chem 266: 5876–5883 - PubMed
-
- Boston RS, Viitanen PV, Vierling E (1996) Molecular chaperones and protein folding in plants. Plant Mol Biol 32: 191–222 - PubMed
-
- Bruce BD, Perry S, Froehlich J, Keegstra K (1994) In vitro import of proteins into chloroplasts. In SB Gelvin, RB Schilperoort, eds, Plant Molecular Biology Manual, Vol J1. Kluwer Academic Publishers, Boston, pp 1–15
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

