Ligand-specific activation of Escherichia coli chemoreceptor transmethylation
- PMID: 15516567
- PMCID: PMC524905
- DOI: 10.1128/JB.186.22.7556-7563.2004
Ligand-specific activation of Escherichia coli chemoreceptor transmethylation
Erratum in
- J Bacteriol. 2005 Jan;187(2):811
Abstract
Adaptation in the chemosensory pathways of bacteria like Escherichia coli is mediated by the enzyme-catalyzed methylation (and demethylation) of glutamate residues in the signaling domains of methyl-accepting chemotaxis proteins (MCPs). MCPs can be methylated in trans, where the methyltransferase (CheR) molecule catalyzing methyl group transfer is tethered to the C terminus of a neighboring receptor. Here, it was shown that E. coli cells exhibited adaptation to attractant stimuli mediated through either engineered or naturally occurring MCPs that were unable to tether CheR as long as another MCP capable of tethering CheR was also present, e.g., either the full-length aspartate or serine receptor (Tar or Tsr). Methylation of isolated membrane samples in which engineered tethering and substrate receptors were coexpressed demonstrated that the truncated substrate receptors (trTsr) were efficiently methylated in the presence of tethering receptors (Tar with methylation sites blocked) relative to samples in which none of the MCPs had tethering sites. The effects of ligand binding on methylation were investigated, and an increase in rate was produced only with serine (the ligand specific for the substrate receptor trTsr); no significant change in rate was produced by aspartate (the ligand specific for the tethering receptor Tar). Although the overall efficiency of methylation was lower, receptor-specific effects were also observed in trTar- and trTsr-containing samples, where neither Tar nor Tsr possessed the CheR binding site at the C terminus. Altogether, the results are consistent with a ligand-induced conformational change that is limited to the methylated receptor dimer and does not spread to adjacent receptor dimers.
Figures


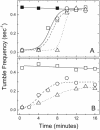

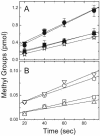


 ) are shown bound to the dimer interface.
) are shown bound to the dimer interface.References
-
- Berg, H. C., and D. A. Brown. 1972. Chemotaxis in Escherichia coli analysed by three-dimensional tracking. Nature 239:500-504. - PubMed
-
- Biemann, H. P., and D. E. Koshland, Jr. 1994. Aspartate receptors of Escherichia coli and Salmonella typhimurium bind ligand with negative and half-of-the-sites cooperativity. Biochemistry 33:629-634. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Miscellaneous

