Phylogenetic characterization of methanogenic assemblages in eutrophic and oligotrophic areas of the Florida Everglades
- PMID: 15528519
- PMCID: PMC525246
- DOI: 10.1128/AEM.70.11.6559-6568.2004
Phylogenetic characterization of methanogenic assemblages in eutrophic and oligotrophic areas of the Florida Everglades
Abstract
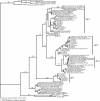
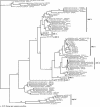
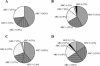
Agricultural activities have produced well-documented changes in the Florida Everglades, including establishment of a gradient in phosphorus concentrations in Water Conservation Area 2A (WCA-2A) of the northern Everglades. An effect of increased phosphorus concentrations is increased methanogenesis in the eutrophic regions compared to the oligotrophic regions of WCA-2A. The goal of this study was to identify relationships between eutrophication and composition and activity of methanogenic assemblages in WCA-2A soils. Distributions of two genes associated with methanogens were characterized in soils taken from WCA-2A: the archaeal 16S rRNA gene and the methyl coenzyme M reductase gene. The richness of methanogen phylotypes was greater in eutrophic than in oligotrophic sites, and sequences related to previously cultivated and uncultivated methanogens were found. A preferential selection for the order Methanomicrobiales was observed in mcrA clone libraries, suggesting primer bias for this group. A greater diversity within the Methanomicrobiales was observed in mcrA clone libraries than in 16S rRNA gene libraries. 16S rRNA phylogenetic analyses revealed a dominance of clones related to Methanosaeta spp., an acetoclastic methanogen dominant in environments with low acetate concentrations. A significant number of clones were related to Methanomicrobiales, an order characterized by species utilizing hydrogen and formate as methanogenic substrates. No representatives of the orders Methanobacteriales and Methanococcales were found in any 16S rRNA clone library, although some Methanobacteriales were found in mcrA libraries. Hydrogenotrophs are the dominant methanogens in WCA-2A, and acetoclastic methanogen genotypes that proliferate in low acetate concentrations outnumber those that typically dominate in higher acetate concentrations.
Figures


References
-
- Bidle, K. A., M. Kastner, and D. H. Bartlett. 1999. A phylogenetic analysis of microbial communities associated with methane hydrate containing marine fluids and sediments in the Cascadia margin (ODP site 892B). FEMS Microbiol. Lett. 177:101-108. - PubMed
-
- Burggraf, S., K. O. Stetter, P. Rouviere, and C. R. Woese. 1991. Methanopyrus kandleri: an archaeal methanogen unrelated to all other known methanogens. Syst. Appl. Microbiol. 14:346-351. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Miscellaneous

