Changes in bacterioplankton composition under different phytoplankton regimens
- PMID: 15528542
- PMCID: PMC525254
- DOI: 10.1128/AEM.70.11.6753-6766.2004
Changes in bacterioplankton composition under different phytoplankton regimens
Abstract
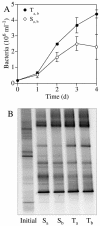
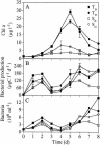
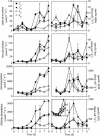
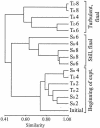
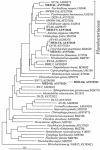
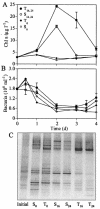
The results of empirical studies have revealed links between phytoplankton and bacterioplankton, such as the frequent correlation between chlorophyll a and bulk bacterial abundance and production. Nevertheless, little is known about possible links at the level of specific taxonomic groups. To investigate this issue, seawater microcosm experiments were performed in the northwestern Mediterranean Sea. Turbulence was used as a noninvasive means to induce phytoplankton blooms dominated by different algae. Microcosms exposed to turbulence became dominated by diatoms, while small phytoflagellates gained importance under still conditions. Denaturing gradient gel electrophoresis (DGGE) of 16S rRNA gene fragments showed that changes in phytoplankton community composition were followed by shifts in bacterioplankton community composition, both as changes in the presence or absence of distinct bacterial phylotypes and as differences in the relative abundance of ubiquitous phylotypes. Sequencing of DGGE bands showed that four Roseobacter phylotypes were present in all microcosms. The microcosms with a higher proportion of phytoflagellates were characterized by four phylotypes of the Bacteroidetes phylum: two affiliated with the family Cryomorphaceae and two with the family Flavobacteriaceae. Two other Flavobacteriaceae phylotypes were characteristic of the diatom-dominated microcosms, together with one Alphaproteobacteria phylotype (Roseobacter) and one Gammaproteobacteria phylotype (Methylophaga). Phylogenetic analyses of published Bacteroidetes 16S rRNA gene sequences confirmed that members of the Flavobacteriaceae are remarkably responsive to phytoplankton blooms, indicating these bacteria could be particularly important in the processing of organic matter during such events. Our data suggest that quantitative and qualitative differences in phytoplankton species composition may lead to pronounced differences in bacterioplankton species composition.
Figures

References
-
- Arin, L., C. Marrasé, M. Maar, F. Peters, M.-M. Sala, and M. Alcaraz. 2002. Combined effects of nutrients and small-scale turbulence in a microcosm experiment. I. Dynamics and size of osmotrophic plankton. Aquat. Microb. Ecol. 29:51-61.
-
- Bergstedt, M., M. M. Hondzo, and J. B. Cotner. 2004. Effects of small scale fluid motion on bacterial growth and respiration. Freshwater Biol. 49:28-40.
-
- Berman, T., and B. Shteinman. 1998. Phytoplankton development and turbulent mixing in Lake Kinneret (1992-1996). J. Plankton Res. 20:709-726.
-
- Bernardet, J.-F., Y. Nakagawa, and B. Holmes. 2002. Proposed minimal standards for describing new taxa of the family Flavobacteriaceae and emended description of the family. Int. J. Syst. Evol. Microbiol. 52:1049-1070. - PubMed
-
- Bidle, K. D., and F. Azam. 2001. Bacterial control of silicon regeneration from diatom detritus: significance of bacterial ectohydrolases and species identity. Limnol. Oceanogr. 46:1606-1623.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

