Genetic recombination between two genotypes of genogroup III bovine noroviruses (BoNVs) and capsid sequence diversity among BoNVs and Nebraska-like bovine enteric caliciviruses
- PMID: 15528717
- PMCID: PMC525163
- DOI: 10.1128/JCM.42.11.5214-5224.2004
Genetic recombination between two genotypes of genogroup III bovine noroviruses (BoNVs) and capsid sequence diversity among BoNVs and Nebraska-like bovine enteric caliciviruses
Abstract
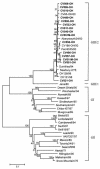
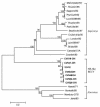
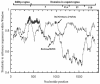
To determine the genogroups and genotypes of bovine enteric caliciviruses (BECVs) circulating in calves, we determined the complete capsid gene sequences of 21 BECVs. The nucleotide and predicted amino acid sequences were compared phylogenetically with those of known human and animal enteric caliciviruses. Based on these analyses, 15 BECVs belonged to Norovirus genogroup III and genotype 2 (GIII/2) and were genetically distinct from human Norovirus GI and GII. Six BECVs had capsid gene sequences similar to that of the unclassified Nebraska (NB)-like BECV. The 15 bovine noroviruses (BoNVs) were more closely related to Bo/NLV/Newbury-2/76/UK (GIII/2) and other known genotype 2 BoNVs than to genotype 1 Bo/NLV/Jena/80/DE. The BoNV Bo/CV521-OH/02/US showed high nucleotide and amino acid identities (84 and 94%, respectively) with the capsid gene of Bo/NLV/Newbury-2/76/UK, whereas the nucleotide and amino acid sequences of the RNA polymerase gene were more closely related to those of Bo/NLV/Jena/80/DE (77 and 87% identities, respectively) than to those of Bo/NLV/Newbury-2/76/UK (69 and 69% identities, respectively), suggesting that Bo/CV521-OH/02/US is a genotype 1-2 recombinant. Gene conversion analysis by the recombinant identification program and SimPlot also predicted that Bo/CV521-OH/02/US was a recombinant. Six NB-like BECVs shared 88 to 92% nucleotide and 94 to 99.5% amino acid identities with the NB BECV in the capsid gene. The results of this study demonstrate genetic diversity in the capsid genes of BECVs circulating in Ohio veal calves, provide new data for coinfections with distinct BECV genotypes or genogroups, and describe the first natural BoNV genotype 1-2 recombinant, analogous to the previously reported human norovirus recombinants.
Figures


References
-
- Ando, T., J. S. Noel, and R. L. Fankhauser. 2000. Genetic classification of “Norwalk-like viruses.” J. Infect. Dis. 181(Suppl. 2):S336-S348. - PubMed
-
- Berke, T., B. Golding, X. Jiang, D. W. Cubitt, M. Wolfaardt, A. W. Smith, and D. O. Matson. 1997. Phylogenetic analysis of the caliciviruses. J. Med. Virol. 52:419-424. - PubMed
-
- Clarke, I. N., and P. R. Lambden. 1997. The molecular biology of caliciviruses. J. Gen. Virol. 78:291-301. - PubMed
-
- Dastjerdi, A. M., J. Green, C. I. Gallimore, D. W. Brown, and J. C. Bridger. 1999. The bovine Newbury agent-2 is genetically more closely related to human SRSVs than to animal caliciviruses. Virology 254:1-5. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

