Yeast chromatin assembly complex 1 protein excludes nonacetylatable forms of histone H4 from chromatin and the nucleus
- PMID: 15542829
- PMCID: PMC529027
- DOI: 10.1128/MCB.24.23.10180-10192.2004
Yeast chromatin assembly complex 1 protein excludes nonacetylatable forms of histone H4 from chromatin and the nucleus
Abstract
In yeast, the establishment and maintenance of a transcriptionally silent chromatin state are dependent upon the acetylation state of the N terminus of histone proteins. Histone H4 proteins that contain mutations in N-terminal lysines disrupt heterochromatin and result in yeast that cannot mate. Introduction of a wild-type copy of histone H4 restores mating, despite the presence of the mutant protein, suggesting that mutant H4 protein is either excluded from, or tolerated in, chromatin. To understand how the cell differentiates wild-type histone and mutant histone in which the four N-terminal lysines were replaced with alanine (H4-4A), we analyzed silencing, growth phenotypes, and the histone composition of chromatin in yeast strains coexpressing equal amounts of wild-type and mutant H4 proteins (histone H4 heterozygote). We found that histone H4 heterozygotes have defects in heterochromatin silencing and growth, implying that mutations in H4 are not completely recessive. Nuclear preparations from histone H4 heterozygotes contained less mutant H4 than wild-type H4, consistent with the idea that cells exclude some of the mutant histone. Surprisingly, the N-terminal nuclear localization signal of H4-4A fused to green fluorescent protein was defective in nuclear localization, while a mutant in which the four lysines were replaced with arginine (H4-4R) appeared to have normal nuclear import, implying a role for the charged state of the acetylatable lysines in the nuclear import of histones. The biased partial exclusion of H4-4A was dependent upon Cac1p, the largest subunit of yeast chromatin assembly factor 1 (CAF-1), as well as upon the karyopherin Kap123p, but was independent of Cac2p, another CAF-1 component, and other chromatin assembly proteins (Hir3p, Nap1p, and Asf1p). We conclude that N-terminal lysines of histone H4 are important for efficient histone nuclear import. In addition, our data support a model whereby Cac1p and Kap123 cooperate to ensure that only appropriately acetylated histone H4 proteins are imported into the nucleus.
Figures





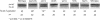

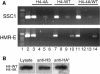
Similar articles
-
The silencing complex SAS-I links histone acetylation to the assembly of repressed chromatin by CAF-I and Asf1 in Saccharomyces cerevisiae.Genes Dev. 2001 Dec 1;15(23):3169-82. doi: 10.1101/gad.929001. Genes Dev. 2001. PMID: 11731480 Free PMC article.
-
Mutational analysis of H3 and H4 N termini reveals distinct roles in nuclear import.J Biol Chem. 2007 Jul 13;282(28):20142-50. doi: 10.1074/jbc.M701989200. Epub 2007 May 15. J Biol Chem. 2007. PMID: 17507373
-
Chromatin assembly factor I mutants defective for PCNA binding require Asf1/Hir proteins for silencing.Mol Cell Biol. 2002 Jan;22(2):614-25. doi: 10.1128/MCB.22.2.614-625.2002. Mol Cell Biol. 2002. PMID: 11756556 Free PMC article.
-
Nuclear import of histones.Biochem Soc Trans. 2020 Dec 18;48(6):2753-2767. doi: 10.1042/BST20200572. Biochem Soc Trans. 2020. PMID: 33300986 Free PMC article. Review.
-
Histone modifications and nuclear architecture: a review.J Histochem Cytochem. 2008 Aug;56(8):711-21. doi: 10.1369/jhc.2008.951251. Epub 2008 May 12. J Histochem Cytochem. 2008. PMID: 18474937 Free PMC article. Review.
Cited by
-
Neocentromeres Provide Chromosome Segregation Accuracy and Centromere Clustering to Multiple Loci along a Candida albicans Chromosome.PLoS Genet. 2016 Sep 23;12(9):e1006317. doi: 10.1371/journal.pgen.1006317. eCollection 2016 Sep. PLoS Genet. 2016. PMID: 27662467 Free PMC article.
-
Molecular functions of the histone acetyltransferase chaperone complex Rtt109-Vps75.Nat Struct Mol Biol. 2008 Sep;15(9):948-56. doi: 10.1038/nsmb.1459. Nat Struct Mol Biol. 2008. PMID: 19172748 Free PMC article.
-
Contribution of CAF-I to anaphase-promoting-complex-mediated mitotic chromatin assembly in Saccharomyces cerevisiae.Eukaryot Cell. 2005 Apr;4(4):673-84. doi: 10.1128/EC.4.4.673-684.2005. Eukaryot Cell. 2005. PMID: 15821127 Free PMC article.
-
A Novel Histone Crosstalk Pathway Important for Regulation of UV-Induced DNA Damage Repair in Saccharomyces cerevisiae.Genetics. 2017 Jul;206(3):1389-1402. doi: 10.1534/genetics.116.195735. Epub 2017 May 18. Genetics. 2017. PMID: 28522541 Free PMC article.
-
Schizosaccharomyces pombe Hat1 (Kat1) is associated with Mis16 and is required for telomeric silencing.Eukaryot Cell. 2012 Sep;11(9):1095-103. doi: 10.1128/EC.00123-12. Epub 2012 Jul 6. Eukaryot Cell. 2012. PMID: 22771823 Free PMC article.
References
-
- Adams, C. R., and R. T. Kamakaka. 1999. Chromatin assembly: biochemical identities and genetic redundancy. Curr. Opin. Genet. Dev. 9:185-190. - PubMed
-
- Ai, X., and M. R. Parthun. 2004. The nuclear Hat1p/Hat2p complex: a molecular link between type B histone acetyltransferases and chromatin assembly. Mol. Cell 14:195-205. - PubMed
-
- Aparicio, O. M., B. L. Billington, and D. E. Gottschling. 1991. Modifiers of position effect are shared between telomeric and silent mating-type loci in S. cerevisiae. Cell 66:1279-1287. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
