An active transposable element, Herves, from the African malaria mosquito Anopheles gambiae
- PMID: 15545643
- PMCID: PMC1449121
- DOI: 10.1534/genetics.104.036145
An active transposable element, Herves, from the African malaria mosquito Anopheles gambiae
Abstract
Transposable elements have proven to be invaluable tools for genetically manipulating a wide variety of plants, animals, and microbes. Some have suggested that they could be used to spread desirable genes, such as refractoriness to Plasmodium infection, through target populations of Anopheles gambiae, thereby disabling the mosquito's ability to transmit malaria. To achieve this, a transposon must remain mobile and intact after the initial introduction into the genome. Endogenous, active class II transposable elements from An. gambiae have not been exploited as gene vectors/drivers because none have been isolated. We report the discovery of an active class II transposable element, Herves, from the mosquito An. gambiae. Herves is a member of a distinct subfamily of hAT elements that includes the hopper-we element from Bactrocera dorsalis and B. cucurbitae. Herves was transpositionally active in mobility assays performed in Drosophila melanogaster S2 cells and developing embryos and was used as a germ-line transformation vector in D. melanogaster. Herves displays an altered target-site preference from the distantly related hAT elements, Hermes and hobo. Herves is also present in An. arabiensis and An. merus with copy numbers similar to that found in An. gambiae. Preliminary data from an East African population are consistent with the element being transpositionally active in mosquitoes.
Figures


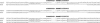

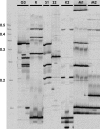
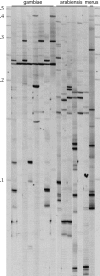
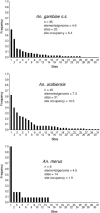
References
-
- Allen, M. L., D. A. O'Brochta, P. W. Atkinson and C. S. LeVesque, 2001. Stable germ-line transformation of Culex quinquefasciatus (Diptera: Culicidae). J. Med. Entomol. 38: 701–710. - PubMed
-
- Altschul, S. F., W. Gish, W. Miller, E. W. Myers and D. J. Lipman, 1990. Basic local alignment search tool. J. Mol. Biol. 215: 403–410. - PubMed
-
- Anxolabehere, D., M. G. Kidwell and G. Periquet, 1988. Molecular characteristics of diverse populations are consistent with the hypothesis of a recent invasion of D. melanogaster by mobile P elements. Mol. Biol. Evol. 5: 252–269. - PubMed
-
- Biessmann, H., M. F. Walter, D. Le, S. Chuan and J. G. Yao, 1999. Moose, a new family of LTR-retrotransposons in the mosquito, Anopheles gambiae. Insect Mol. Biol. 8: 201–212. - PubMed
-
- Calvi, B. R., T. J. Hong, S. D. Findley and W. M. Gelbart, 1991. Evidence for a common evolutionary origin of inverted terminal repeat transposons in Drosophila and plants: hobo, Activator and Tam3. Cell 66: 465–471. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

