Interaction between the oxa1 and rmp1 genes modulates respiratory complex assembly and life span in Podospora anserina
- PMID: 15545650
- PMCID: PMC1449539
- DOI: 10.1534/genetics.104.033837
Interaction between the oxa1 and rmp1 genes modulates respiratory complex assembly and life span in Podospora anserina
Abstract
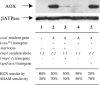
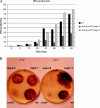
A causal link between deficiency of the cytochrome respiratory pathway and life span was previously shown in the filamentous fungus Podospora anserina. To gain more insight into the relationship between mitochondrial function and life span, we have constructed a strain carrying a thermosensitive mutation of the gene oxa1. OXA1 is a membrane protein conserved from bacteria to human. The mitochondrial OXA1 protein is involved in the assembly/insertion of several respiratory complexes. We show here that oxa1 is an essential gene in P. anserina. The oxa1(ts) mutant exhibits severe defects in the respiratory complexes I and IV, which are correlated with an increased life span, a strong induction of the alternative oxidase, and a reduction in ROS production. However, there is no causal link between alternative oxidase level and life span. We also show that in the oxa1(ts) mutant, the extent of the defects in complexes I and IV and the life-span increase depends on the essential gene rmp1. The RMP1 protein, whose function is still unknown, can be localized in the mitochondria and/or the cytosolic compartment, depending on the developmental stage. We propose that the RMP1 protein could be involved in the process of OXA1-dependent protein insertion.
Figures


References
-
- Albert, B., and C. H. Sellem, 2002. Dynamics of the mitochondrial genome during Podospora anserina aging. Curr. Genet. 40: 365–373. - PubMed
-
- Altamura, N., N. Capitanio, N. Bonnefoy, S. Papa and G. Dujardin, 1996. The Saccharomyces cerevisiae OXA1 gene is required for the correct assembly of cytochrome c oxidase and oligomycin-sensitive ATP synthase. FEBS Lett. 382: 111–115. - PubMed
-
- Barreau, C., C. Sellem, P. Silar, A. Sainsard-Chanet and B. Turcq, 2002. A rapid and efficient method using chromoslots to assign any newly cloned DNA sequence to its cognate chromosome in the filamentous fungus Podospora anserina. FEMS Microbiol. Lett. 216: 55–60. - PubMed
-
- Bauer, M., M. Behrens, K. Esser, G. Michaelis and E. Pratje, 1994. PET1402, a nuclear gene required for proteolytic processing of cytochrome oxidase subunit 2 in yeast. Mol. Gen. Genet. 245: 272–278. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
LinkOut - more resources
Full Text Sources

