Analysis of mutant phenotypes and splicing defects demonstrates functional collaboration between the large and small subunits of the essential splicing factor U2AF in vivo
- PMID: 15548596
- PMCID: PMC545896
- DOI: 10.1091/mbc.e04-09-0768
Analysis of mutant phenotypes and splicing defects demonstrates functional collaboration between the large and small subunits of the essential splicing factor U2AF in vivo
Abstract
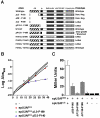
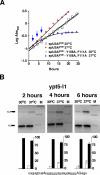
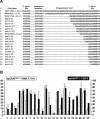
The heterodimeric splicing factor U2AF plays an important role in 3' splice site selection, but the division of labor between the two subunits in vivo remains unclear. In vitro assays led to the proposal that the human large subunit recognizes 3' splice sites with extensive polypyrimidine tracts independently of the small subunit. We report in vivo analysis demonstrating that all five domains of spU2AFLG are essential for viability; a partial deletion of the linker region, which forms the small subunit interface, produces a severe growth defect and an aberrant morphology. A small subunit zinc-binding domain mutant confers a similar phenotype, suggesting that the heterodimer functions as a unit during splicing in Schizosaccharomyces pombe. As this is not predicted by the model for metazoan 3' splice site recognition, we sought introns for which the spU2AFLG and spU2AFSM make distinct contributions by analyzing diverse splicing events in strains harboring mutations in each partner. Requirements for the two subunits are generally parallel and, moreover, do not correlate with the length or strength of the 3' pyrimidine tract. These and other studies performed in fission yeast support a model for 3' splice site recognition in which the two subunits of U2AF functionally collaborate in vivo.
Figures



References
-
- Abovich, N., Liao, X., and Rosbash, M. (1994). The yeast MUD2 protein: an interaction with PRP11 defines a bridge between commitment complex and U2 snRNP addition. Genes Dev. 8, 843-854. - PubMed
-
- Beales, M., Flay, N., McKinney, R., Habara, Y., Ohshima, Y., Tani, T., and Potashkin, J. (2000). Mutations in the large subunit of U2AF disrupt pre-mRNA splicing, cell cycle progression and nuclear structure. Yeast 16, 1001-1013. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

