Long-range control of gene expression: emerging mechanisms and disruption in disease
- PMID: 15549674
- PMCID: PMC1196435
- DOI: 10.1086/426833
Long-range control of gene expression: emerging mechanisms and disruption in disease
Abstract
Transcriptional control is a major mechanism for regulating gene expression. The complex machinery required to effect this control is still emerging from functional and evolutionary analysis of genomic architecture. In addition to the promoter, many other regulatory elements are required for spatiotemporally and quantitatively correct gene expression. Enhancer and repressor elements may reside in introns or up- and downstream of the transcription unit. For some genes with highly complex expression patterns--often those that function as key developmental control genes--the cis-regulatory domain can extend long distances outside the transcription unit. Some of the earliest hints of this came from disease-associated chromosomal breaks positioned well outside the relevant gene. With the availability of wide-ranging genome sequence comparisons, strong conservation of many noncoding regions became obvious. Functional studies have shown many of these conserved sites to be transcriptional regulatory elements that sometimes reside inside unrelated neighboring genes. Such sequence-conserved elements generally harbor sites for tissue-specific DNA-binding proteins. Developmentally variable chromatin conformation can control protein access to these sites and can regulate transcription. Disruption of these finely tuned mechanisms can cause disease. Some regulatory element mutations will be associated with phenotypes distinct from any identified for coding-region mutations.
Figures

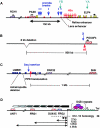

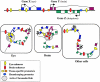
References
Electronic-Database Information
-
- Online Mendelian Inheritance in Man (OMIM), http://www.ncbi.nlm.nih.gov/Omim/ (for aniridia, deafness type 3, Rieger syndrome type 1, Greig cephalopolysyndactyly syndrome, MAF, LD, blepharophimosis-ptosis-epicanthus inversus syndrome, CD, HPE2, HPE3, PPD, SHFM1, FSHD, α-thalassemia, CDLS, and lactase expression)
References
-
- Bakker E, Wijmenga C, Vossen RH, Padberg GW, Hewitt J, van der WM, Rasmussen K, Frants RR (1995) The FSHD-linked locus D4F104S1 (p13E-11) on 4q35 has a homologue on 10qter. Muscle Nerve 2:S39–S44 - PubMed
-
- Belloni E, Muenke M, Roessler E, Traverso G, Siegel-Bartelt J, Frumkin A, Mitchell HF, Donis-Keller H, Helms C, Hing AV, Heng HH, Koop B, Martindale D, Rommens JM, Tsui LC, Scherer SW (1996) Identification of Sonic hedgehog as a candidate gene responsible for holoprosencephaly. Nat Genet 14:353–356 10.1038/ng1196-353 - DOI - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical

