HMW1 is required for stability and localization of HMW2 to the attachment organelle of Mycoplasma pneumoniae
- PMID: 15576770
- PMCID: PMC532429
- DOI: 10.1128/JB.186.24.8221-8228.2004
HMW1 is required for stability and localization of HMW2 to the attachment organelle of Mycoplasma pneumoniae
Abstract
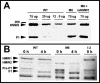
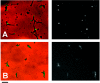
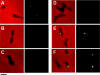
The cytoskeletal proteins HMW1 and HMW2 are components of the terminal organelle of the cell wall-less bacterium Mycoplasma pneumoniae. HMW1 is required for a tapered, filamentous morphology but exhibits accelerated turnover in the absence of HMW2. Here, we report that a reciprocal dependency exists between HMW1 and HMW2, with HMW2 subject to accelerated turnover with the loss of HMW1. Furthermore, the instability of HMW2 correlated with its failure to localize to the attachment organelle. The C-terminal domain of HMW1 is essential for both function and its accelerated turnover in the absence of HMW2. We constructed HMW1 deletion derivatives lacking portions of this domain and examined each for stability and function. The C-terminal 41 residues were particularly important for proper localization and function in cell morphology and P1 localization, but the entire C-terminal domain was required to stabilize HMW2. The significance of these findings in the context of attachment organelle assembly is considered.
Figures





Similar articles
-
Deletion analysis identifies key functional domains of the cytadherence-associated protein HMW2 of Mycoplasma pneumoniae.Mol Microbiol. 2003 Dec;50(5):1507-16. doi: 10.1046/j.1365-2958.2003.03807.x. Mol Microbiol. 2003. PMID: 14651634
-
Loss of HMW1 and HMW3 in noncytadhering mutants of Mycoplasma pneumoniae occurs post-translationally.Proc Natl Acad Sci U S A. 1997 Dec 9;94(25):13979-84. doi: 10.1073/pnas.94.25.13979. Proc Natl Acad Sci U S A. 1997. PMID: 9391138 Free PMC article.
-
Stability of Mycoplasma pneumoniae cytadherence-accessory protein HMW1 correlates with its association with the triton shell.J Bacteriol. 2001 Jun;183(12):3680-8. doi: 10.1128/JB.183.12.3680-3688.2001. J Bacteriol. 2001. PMID: 11371532 Free PMC article.
-
Structure, function, and assembly of the terminal organelle of Mycoplasma pneumoniae.FEMS Microbiol Lett. 2001 Apr 20;198(1):1-7. doi: 10.1111/j.1574-6968.2001.tb10610.x. FEMS Microbiol Lett. 2001. PMID: 11325545 Review.
-
Mycoplasma pneumoniae cytadherence: organization and assembly of the attachment organelle.Trends Microbiol. 1998 Jan;6(1):15-8. doi: 10.1016/S0966-842X(97)01168-2. Trends Microbiol. 1998. PMID: 9481818 Review.
Cited by
-
Synthesis, stability, and function of cytadhesin P1 and accessory protein B/C complex of Mycoplasma pneumoniae.J Bacteriol. 2006 Jan;188(2):569-75. doi: 10.1128/JB.188.2.569-575.2006. J Bacteriol. 2006. PMID: 16385047 Free PMC article.
-
Insights into the pathogenesis of Mycoplasma pneumoniae (Review).Mol Med Rep. 2016 Nov;14(5):4030-4036. doi: 10.3892/mmr.2016.5765. Epub 2016 Sep 23. Mol Med Rep. 2016. PMID: 27667580 Free PMC article. Review.
-
Periodicity in Attachment Organelle Revealed by Electron Cryotomography Suggests Conformational Changes in Gliding Mechanism of Mycoplasma pneumoniae.mBio. 2016 Apr 12;7(2):e00243-16. doi: 10.1128/mBio.00243-16. mBio. 2016. PMID: 27073090 Free PMC article.
-
The stability of cytadherence proteins in Mycoplasma pneumoniae requires activity of the protein kinase PrkC.Infect Immun. 2010 Jan;78(1):184-92. doi: 10.1128/IAI.00958-09. Epub 2009 Oct 26. Infect Immun. 2010. PMID: 19858294 Free PMC article.
-
Functional analysis of the Mycoplasma genitalium MG312 protein reveals a specific requirement of the MG312 N-terminal domain for gliding motility.J Bacteriol. 2007 Oct;189(19):7014-23. doi: 10.1128/JB.00975-07. Epub 2007 Aug 3. J Bacteriol. 2007. PMID: 17675381 Free PMC article.
References
-
- Akiyama, Y., A. Kihara, and K. Ito. 1996. Subunit a of proton ATPase F0 sector is a substrate of the FtsH protease in Escherichia coli. FEBS Lett. 399:26-28. - PubMed
-
- Balish, M. F., and D. C. Krause. 2002. Cytadherence and the cytoskeleton, p. 491-518. In S. Razin and R. Herrmann (ed.), Molecular biology and pathogenicity of the mycoplasmas. Kluwer Academic/Plenum Publishers, New York, N.Y.
-
- Balish, M. F., S. M. Ross, M. Fisseha, and D. C. Krause. 2003. Deletion analysis identifies key functional domains of the cytadherence-associated protein HMW2 of Mycoplasma pneumoniae. Mol. Microbiol. 50:1507-1516. - PubMed
-
- Balish, M. F., R. T. Santurri, A. M. Ricci, K. K. Lee, and D. C. Krause. 2003. Localization of Mycoplasma pneumoniae cytadherence-associated protein HMW2 by fusion with green fluorescent protein: implications for attachment organelle structure. Mol. Microbiol. 47:49-60. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

