Bacillus subtilis operon encoding a membrane receptor for bacteriophage SPP1
- PMID: 15576783
- PMCID: PMC532427
- DOI: 10.1128/JB.186.24.8337-8346.2004
Bacillus subtilis operon encoding a membrane receptor for bacteriophage SPP1
Abstract
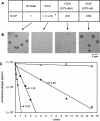
The results reported here have identified yueB as the essential gene involved in irreversible binding of bacteriophage SPP1 to Bacillus subtilis. First, a deletion in an SPP1-resistant (pha-2) strain, covering most of the yueB gene, could be complemented by a xylose-inducible copy of yueB inserted at amyE. Second, disruption of yueB by insertion of a pMutin4 derivative resulted in a phage resistance phenotype regardless of the presence or absence of IPTG (isopropyl-beta-D-thiogalactopyranoside). YueB homologues are widely distributed in gram-positive bacteria. The protein Pip, which also serves as a phage receptor in Lactococcus lactis, belongs to the same family. yueB encodes a membrane protein of approximately 120 kDa, detected in immunoblots together with smaller forms that may be processed products arising from cleavage of its long extracellular domain. Insertional inactivation of yueB and the surrounding genes indicated that yueB is part of an operon which includes at least the upstream genes yukE, yukD, yukC, and yukBA. Disruption of each of the genes in the operon allowed efficient irreversible adsorption, provided that yueB expression was retained. Under these conditions, however, smaller plaques were produced, a phenotype which was particularly noticeable in yukE mutant strains. Interestingly, such reduction in plaque size was not correlated with a decreased adsorption rate. Overall, these results provide the first demonstration of a membrane-bound protein acting as a phage receptor in B. subtilis and suggest an additional involvement of the yukE operon in a step subsequent to irreversible adsorption.
Figures






Similar articles
-
Phage SPP1 reversible adsorption to Bacillus subtilis cell wall teichoic acids accelerates virus recognition of membrane receptor YueB.J Bacteriol. 2008 Jul;190(14):4989-96. doi: 10.1128/JB.00349-08. Epub 2008 May 16. J Bacteriol. 2008. PMID: 18487323 Free PMC article.
-
The ectodomain of the viral receptor YueB forms a fiber that triggers ejection of bacteriophage SPP1 DNA.J Biol Chem. 2006 Apr 28;281(17):11464-70. doi: 10.1074/jbc.M513625200. Epub 2006 Feb 14. J Biol Chem. 2006. PMID: 16481324
-
Role of bacteriophage SPP1 tail spike protein gp21 on host cell receptor binding and trigger of phage DNA ejection.Mol Microbiol. 2012 Jan;83(2):289-303. doi: 10.1111/j.1365-2958.2011.07931.x. Epub 2011 Dec 15. Mol Microbiol. 2012. PMID: 22171743
-
Membrane Penetration by Bacterial Viruses.J Virol. 2017 Jun 9;91(13):e00162-17. doi: 10.1128/JVI.00162-17. Print 2017 Jul 1. J Virol. 2017. PMID: 28404851 Free PMC article. Review.
-
[Gluconate operon of Bacillus subtilis: a model for enzyme induction of gram-positive bacteria].Seikagaku. 1988 Jun;60(6):436-41. Seikagaku. 1988. PMID: 3150421 Review. Japanese. No abstract available.
Cited by
-
Molecular Basis for Lytic Bacteriophage Resistance in Enterococci.mBio. 2016 Aug 30;7(4):e01304-16. doi: 10.1128/mBio.01304-16. mBio. 2016. PMID: 27578757 Free PMC article.
-
Symbiosis of a lytic bacteriophage and Yersinia pestis and characteristics of plague in Marmota himalayana.Appl Environ Microbiol. 2024 Aug 21;90(8):e0099524. doi: 10.1128/aem.00995-24. Epub 2024 Jul 18. Appl Environ Microbiol. 2024. PMID: 39023266 Free PMC article.
-
Signatures of selection in core and accessory genomes indicate different ecological drivers of diversification among Bacillus cereus clades.Mol Ecol. 2022 Jul;31(13):3584-3597. doi: 10.1111/mec.16490. Epub 2022 May 17. Mol Ecol. 2022. PMID: 35510788 Free PMC article.
-
Genetic Signatures of Dairy Lactobacillus casei Group.Front Microbiol. 2018 Oct 30;9:2611. doi: 10.3389/fmicb.2018.02611. eCollection 2018. Front Microbiol. 2018. PMID: 30425707 Free PMC article.
-
Bacteria elicit a phage tolerance response subsequent to infection of their neighbors.EMBO J. 2022 Feb 1;41(3):e109247. doi: 10.15252/embj.2021109247. Epub 2021 Dec 8. EMBO J. 2022. PMID: 34878184 Free PMC article.
References
-
- Adams, M. H. 1959. Bacteriophages. Wiley Interscience, New York, N.Y.
-
- Biswal, N., A. K. Kleinschmidt, H. C. Spatz, and T. A. Trautner. 1967. Physical properties of the DNA of bacteriophage SP50. Mol. Gen. Genet. 100:39-55. - PubMed
-
- Casadaban, M. J., and S. N. Cohen. 1980. Analysis of gene control signals by DNA fusion and cloning in Escherichia coli. J. Mol. Biol. 138:179-207. - PubMed
-
- Errington, J., J. Bath, and L. J. Wu. 2001. DNA transport in bacteria. Nat. Rev. Mol. Cell Biol. 2:538-545. - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Research Materials
Miscellaneous

