Structure of the complex between the cytosolic chaperonin CCT and phosducin-like protein
- PMID: 15583139
- PMCID: PMC536017
- DOI: 10.1073/pnas.0405070101
Structure of the complex between the cytosolic chaperonin CCT and phosducin-like protein
Abstract
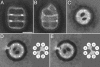
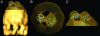
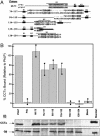
The three-dimensional structure of the complex formed between the cytosolic chaperonin CCT (chaperonin containing TCP-1) and phosducin (Pdc)-like protein (PhLP), a regulator of CCT activity, has been solved by cryoelectron microscopy. Binding of PhLP to CCT occurs through only one of the chaperonin rings, and the protein does not occupy the central folding cavity but rather sits above it through interactions with two regions on opposite sides of the ring. This causes the apical domains of the CCT subunits to close in, thus excluding access to the folding cavity. The atomic model of PhLP generated from several atomic structures of the homologous Pdc fits very well with the mass of the complex attributable to PhLP and predicts the involvement of several sequences of PhLP in CCT binding. Binding experiments performed with PhLP/Pdc chimeric proteins, taking advantage of the fact that Pdc does not interact with CCT, confirm that both the N- and C-terminal domains of PhLP are involved in CCT binding and that several regions suggested by the docking experiment are indeed critical in the interaction with the cytosolic chaperonin.
Figures

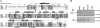

Similar articles
-
Phosducin-like protein regulates G-protein betagamma folding by interaction with tailless complex polypeptide-1alpha: dephosphorylation or splicing of PhLP turns the switch toward regulation of Gbetagamma folding.J Biol Chem. 2005 May 20;280(20):20042-50. doi: 10.1074/jbc.M409233200. Epub 2005 Mar 2. J Biol Chem. 2005. PMID: 15745879
-
Regulatory interaction of phosducin-like protein with the cytosolic chaperonin complex.Proc Natl Acad Sci U S A. 2002 Jun 11;99(12):7962-7. doi: 10.1073/pnas.112075699. Proc Natl Acad Sci U S A. 2002. PMID: 12060742 Free PMC article.
-
Structures of the Gβ-CCT and PhLP1-Gβ-CCT complexes reveal a mechanism for G-protein β-subunit folding and Gβγ dimer assembly.Proc Natl Acad Sci U S A. 2015 Feb 24;112(8):2413-8. doi: 10.1073/pnas.1419595112. Epub 2015 Feb 9. Proc Natl Acad Sci U S A. 2015. PMID: 25675501 Free PMC article.
-
The substrate recognition mechanisms in chaperonins.J Mol Recognit. 2004 Mar-Apr;17(2):85-94. doi: 10.1002/jmr.654. J Mol Recognit. 2004. PMID: 15027029 Review.
-
Structure and function of a protein folding machine: the eukaryotic cytosolic chaperonin CCT.FEBS Lett. 2002 Oct 2;529(1):11-6. doi: 10.1016/s0014-5793(02)03180-0. FEBS Lett. 2002. PMID: 12354605 Review.
Cited by
-
Structural basis of plp2-mediated cytoskeletal protein folding by TRiC/CCT.Sci Adv. 2023 Mar 17;9(11):eade1207. doi: 10.1126/sciadv.ade1207. Epub 2023 Mar 15. Sci Adv. 2023. PMID: 36921056 Free PMC article.
-
The TRiCky Business of Protein Folding in Health and Disease.Front Cell Dev Biol. 2022 May 5;10:906530. doi: 10.3389/fcell.2022.906530. eCollection 2022. Front Cell Dev Biol. 2022. PMID: 35602608 Free PMC article. Review.
-
The molecular architecture of the eukaryotic chaperonin TRiC/CCT.Structure. 2012 May 9;20(5):814-25. doi: 10.1016/j.str.2012.03.007. Epub 2012 Apr 12. Structure. 2012. PMID: 22503819 Free PMC article.
-
Modulation of phosducin-like protein 3 (PhLP3) levels promotes cytoskeletal remodelling in a MAPK and RhoA-dependent manner.PLoS One. 2011;6(12):e28271. doi: 10.1371/journal.pone.0028271. Epub 2011 Dec 9. PLoS One. 2011. PMID: 22174782 Free PMC article.
-
Activities of the chaperonin containing TCP-1 (CCT): implications for cell cycle progression and cytoskeletal organisation.Cell Stress Chaperones. 2009 Jan;14(1):23-31. doi: 10.1007/s12192-008-0057-x. Epub 2008 Jul 2. Cell Stress Chaperones. 2009. PMID: 18595008 Free PMC article. Review.
References
-
- Ellis, R. J. (1996) in The Chaperonins, ed. Ellis, R. J. (Academic, San Diego), pp. 2–25.
-
- Ellis, R. J. & Hartl, F. U. (1999) Curr. Opin. Struct. Biol. 9, 102–110. - PubMed
-
- Gutsche, I., Essen, L. O. & Baumeister, W. (1999) J. Mol. Biol. 293, 295–312. - PubMed
-
- Braig, K., Otwinowski, Z., Hegde, R., Boisvert, D. C., Joachimiak, A., Horwich, A. L. & Sigler, P. B. (1994) Nature 371, 578–586. - PubMed
-
- Ditzel, L., Löwe, J., Stock, D., Stetter, K. O., Huber, H., Huber, R. & Steinbacher, S. (1998) Cell 93, 125–138. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

