Genome sequence of the deep-sea gamma-proteobacterium Idiomarina loihiensis reveals amino acid fermentation as a source of carbon and energy
- PMID: 15596722
- PMCID: PMC539801
- DOI: 10.1073/pnas.0407638102
Genome sequence of the deep-sea gamma-proteobacterium Idiomarina loihiensis reveals amino acid fermentation as a source of carbon and energy
Abstract
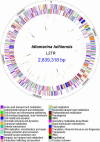
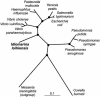
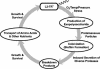
We report the complete genome sequence of the deep-sea gamma-proteobacterium, Idiomarina loihiensis, isolated recently from a hydrothermal vent at 1,300-m depth on the Loihi submarine volcano, Hawaii. The I. loihiensis genome comprises a single chromosome of 2,839,318 base pairs, encoding 2,640 proteins, four rRNA operons, and 56 tRNA genes. A comparison of I. loihiensis to the genomes of other gamma-proteobacteria reveals abundance of amino acid transport and degradation enzymes, but a loss of sugar transport systems and certain enzymes of sugar metabolism. This finding suggests that I. loihiensis relies primarily on amino acid catabolism, rather than on sugar fermentation, for carbon and energy. Enzymes for biosynthesis of purines, pyrimidines, the majority of amino acids, and coenzymes are encoded in the genome, but biosynthetic pathways for Leu, Ile, Val, Thr, and Met are incomplete. Auxotrophy for Val and Thr was confirmed by in vivo experiments. The I. loihiensis genome contains a cluster of 32 genes encoding enzymes for exopolysaccharide and capsular polysaccharide synthesis. It also encodes diverse peptidases, a variety of peptide and amino acid uptake systems, and versatile signal transduction machinery. We propose that the source of amino acids for I. loihiensis growth are the proteinaceous particles present in the deep sea hydrothermal vent waters. I. loihiensis would colonize these particles by using the secreted exopolysaccharide, digest these proteins, and metabolize the resulting peptides and amino acids. In summary, the I. loihiensis genome reveals an integrated mechanism of metabolic adaptation to the constantly changing deep-sea hydrothermal ecosystem.
Figures


Similar articles
-
Complete genome sequence of Crassaminicella sp. 143-21,isolated from a deep-sea hydrothermal vent.Mar Genomics. 2022 Apr;62:100899. doi: 10.1016/j.margen.2021.100899. Epub 2021 Sep 4. Mar Genomics. 2022. PMID: 35246304
-
Idiomarina loihiensis sp. nov., a halophilic gamma-Proteobacterium from the Lō'ihi submarine volcano, Hawai'i.Int J Syst Evol Microbiol. 2003 Nov;53(Pt 6):1873-9. doi: 10.1099/ijs.0.02701-0. Int J Syst Evol Microbiol. 2003. PMID: 14657116
-
Locked chromophore analogs reveal that photoactive yellow protein regulates biofilm formation in the deep sea bacterium Idiomarina loihiensis.J Am Chem Soc. 2009 Dec 2;131(47):17443-51. doi: 10.1021/ja9057103. J Am Chem Soc. 2009. PMID: 19891493
-
Bioinformatic analysis of an unusual gene-enzyme relationship in the arginine biosynthetic pathway among marine gamma proteobacteria: implications concerning the formation of N-acetylated intermediates in prokaryotes.BMC Genomics. 2006 Jan 12;7:4. doi: 10.1186/1471-2164-7-4. BMC Genomics. 2006. PMID: 16409639 Free PMC article.
-
Deep-sea vent epsilon-proteobacterial genomes provide insights into emergence of pathogens.Proc Natl Acad Sci U S A. 2007 Jul 17;104(29):12146-50. doi: 10.1073/pnas.0700687104. Epub 2007 Jul 5. Proc Natl Acad Sci U S A. 2007. PMID: 17615243 Free PMC article.
Cited by
-
Prokaryotic Response to Phytodetritus-Derived Organic Material in Epi- and Mesopelagic Antarctic Waters.Front Microbiol. 2020 Jun 9;11:1242. doi: 10.3389/fmicb.2020.01242. eCollection 2020. Front Microbiol. 2020. PMID: 32582131 Free PMC article.
-
Pan-Genomic and Transcriptomic Analyses of Marine Pseudoalteromonas agarivorans Hao 2018 Revealed Its Genomic and Metabolic Features.Mar Drugs. 2022 Mar 31;20(4):248. doi: 10.3390/md20040248. Mar Drugs. 2022. PMID: 35447921 Free PMC article.
-
The genome of deep-sea vent chemolithoautotroph Thiomicrospira crunogena XCL-2.PLoS Biol. 2006 Nov;4(12):e383. doi: 10.1371/journal.pbio.0040383. PLoS Biol. 2006. PMID: 17105352 Free PMC article.
-
Microbial community transcriptomes reveal microbes and metabolic pathways associated with dissolved organic matter turnover in the sea.Proc Natl Acad Sci U S A. 2010 Sep 21;107(38):16420-7. doi: 10.1073/pnas.1010732107. Epub 2010 Aug 31. Proc Natl Acad Sci U S A. 2010. PMID: 20807744 Free PMC article.
-
Genome analysis of Rubritalea profundi SAORIC-165T, the first deep-sea verrucomicrobial isolate, from the northwestern Pacific Ocean.J Microbiol. 2019 May;57(5):413-422. doi: 10.1007/s12275-019-8712-8. Epub 2019 Feb 26. J Microbiol. 2019. PMID: 30806980
References
-
- Karl, D. M., McMurtry, G. M., Malahoff, A. & Garcia, M. O. (1988) Nature 335, 532–535.
-
- Karl, D. M. (1995) in The Microbiology of Deep-Sea Hydrothermal Vents, ed. Karl, D. M. (CRC Press, Boca Raton, FL), pp. 35–124.
-
- Hilton, D. R., McMurtry, G. M. & Goff, F. (1998) Nature 396, 359–362.
-
- Reysenbach, A. L. & Shock, E. (2002) Science 296, 1077–1082. - PubMed
-
- Nelson, K. E. (2003) Environ. Microbiol. 5, 1223–1225. - PubMed
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
Miscellaneous

