Loss of pollen-S function in two self-compatible selections of Prunus avium is associated with deletion/mutation of an S haplotype-specific F-box gene
- PMID: 15598801
- PMCID: PMC544488
- DOI: 10.1105/tpc.104.026963
Loss of pollen-S function in two self-compatible selections of Prunus avium is associated with deletion/mutation of an S haplotype-specific F-box gene
Abstract
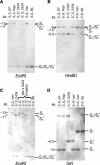
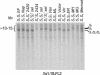
Recently, an S haplotype-specific F-box (SFB) gene has been proposed as a candidate for the pollen-S specificity gene of RNase-mediated gametophytic self-incompatibility in Prunus (Rosaceae). We have examined two pollen-part mutant haplotypes of sweet cherry (Prunus avium). Both were found to retain the S-RNase, which determines stylar specificity, but one (S3' in JI 2434) has a deletion including the haplotype-specific SFB gene, and the other (S4' in JI 2420) has a frame-shift mutation of the haplotype-specific SFB gene, causing amino acid substitutions and premature termination of the protein. The loss or significant alteration of this highly polymorphic gene and the concomitant loss of pollen self-incompatibility function provides compelling evidence that the SFB gene encodes the pollen specificity component of self-incompatibility in Prunus. These loss-of-function mutations are inconsistent with SFB being the inactivator of non-self S-RNases and indicate the presence of a general inactivation mechanism, with SFB conferring specificity by protecting self S-RNases from inactivation.
Figures






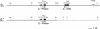

References
-
- Bošković, R., and Tobutt, K.R. (1996). Correlation of stylar ribonuclease zymograms with incompatibility alleles in sweet cherry. Euphytica 90, 245–250.
-
- Bošković, R., and Tobutt, K.R. (1997). Inheritance and linkage relationships of isoenzymes in two interspecific cherry progenies. Euphytica 103, 273–286.
-
- Bošković, R., Russell, K., and Tobutt, K.R. (1997). Inheritance of stylar ribonucleases in cherry progenies, and reassignment of incompatibility alleles to two incompatibility groups. Euphytica 95, 221–228.
-
- Bošković, R., Tobutt, K.R., and Russell, K. (1998). Selection of sweet cherry seedlings homozygous for self-compatibility. Acta Hortic. 484, 249–253.
-
- Bošković, R., Tobutt, K.R., Schmidt, H., and Sonneveld, T. (2000). Re-examination of (in)compatibility genotypes of two John Innes self-compatible sweet cherry selections. Theor. Appl. Genet. 101, 234–240.
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

