Mass spectrometry analysis of Arabidopsis histone H3 reveals distinct combinations of post-translational modifications
- PMID: 15598823
- PMCID: PMC545460
- DOI: 10.1093/nar/gkh992
Mass spectrometry analysis of Arabidopsis histone H3 reveals distinct combinations of post-translational modifications
Abstract
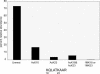
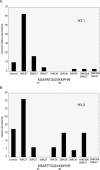
Chromatin is regulated at many different levels, from higher-order packing to individual nucleosome placement. Recent studies have shown that individual histone modifications, and combinations thereof, play a key role in modulating chromatin structure and gene activity. Reported here is an analysis of Arabidopsis histone H3 modifications by nanoflow-HPLC coupled to electrospray ionization on a hybrid linear ion trap-Fourier transform mass spectrometer (LTQ/FTMS). We find that the sites of acetylation and methylation, in general, correlate well with other plants and animals. Two well-studied modifications, dimethylation of Lys-9 (correlated with silencing) and acetylation of Lys-14 (correlated with active chromatin) while abundant by themselves were rarely found on the same histone H3 tail. In contrast, dimethylation at Lys-27 and monomethylation at Lys-36 were commonly found together. Interestingly, acetylation at Lys-9 was found only in a low percentage of histones while acetylation of Lys-14 was very abundant. The two histone H3 variants, H3.1 and H3.2, also differ in the abundance of silencing and activating marks confirming other studies showing that the replication-independent histone H3 is enriched in active chromatin.
Figures



References
-
- Luger K., Mader,A.W., Richmond,R.K., Sargent,D.F. and Richmond,T.J. (1997) Crystal structure of the nucleosome core particle at 2.8 Å resolution. Nature, 389, 251–260. - PubMed
-
- Tagami H., Ray-Gallet,D., Almouzni,G. and Nakatani,Y. (2004) Histone H3.1 and H3.3 complexes mediate nucleosome assembly pathways dependent or independent of DNA synthesis. Cell, 116, 51–61. - PubMed
-
- Tyler J.K., Adams,C.R., Chen,S.R., Kobayashi,R., Kamakaka,R.T. and Kadonaga,J.T. (1999) The RCAF complex mediates chromatin assembly during DNA replication and repair. Nature, 402, 555–560. - PubMed
-
- Ray-Gallet D., Quivy,J.P., Scamps,C., Martini,E.M., Lipinski,M. and Almouzni,G. (2002) HIRA is critical for a nucleosome assembly pathway independent of DNA synthesis. Mol. Cell, 9, 1091–1100. - PubMed
-
- Ahmad K. and Henikoff,S. (2002) The histone variant H3.3 marks active chromatin by replication-independent nucleosome assembly. Mol. Cell, 9, 1191–1200. - PubMed
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

