Pa-AGOG, the founding member of a new family of archaeal 8-oxoguanine DNA-glycosylases
- PMID: 15604455
- PMCID: PMC545463
- DOI: 10.1093/nar/gkh995
Pa-AGOG, the founding member of a new family of archaeal 8-oxoguanine DNA-glycosylases
Abstract
Oxidative damage represents a major threat to genomic stability, as the major product of DNA oxidation, 8-oxoguanine (GO), frequently mispairs with adenine during replication. In order to prevent these mutagenic events, organisms have evolved GO-DNA glycosylases that remove this oxidized base from DNA. We were interested to find out how GO is processed in the hyperthermophilic archaeon Pyrobaculum aerophilum, which lives at temperatures around 100 degrees C. To this end, we searched its genome for open reading frames (ORFs) bearing the principal hallmark of GO-DNA glycosylases: a helix-hairpin-helix motif and a glycine/proline-rich sequence followed by an absolutely conserved aspartate (HhH-GPD motif). Interestingly, although the P.aerophilum genome encodes three such ORFs, none of these encodes the potent GO-processing activity detected in P.aerophilum extracts. Fractionation of the extracts, followed by analysis of the active fractions by denaturing polyacrylamide gel electrophoresis, showed that the GO-processing enzyme has a molecular size of approximately 30 kDa. Mass spectrometric analysis of proteins in this size range identified several peptides originating from P.aerophilum ORF PAE2237. We now show that PAE2237 encodes AGOG (Archaeal GO-Glycosylase), the founding member of a new family of DNA glycosylases, which can remove GO from single- and double-stranded substrates with great efficiency.
Figures





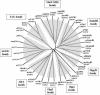
Similar articles
-
Mutational studies of Pa-AGOG DNA glycosylase from the hyperthermophilic crenarchaeon Pyrobaculum aerophilum.DNA Repair (Amst). 2009 Jul 4;8(7):857-64. doi: 10.1016/j.dnarep.2009.03.009. Epub 2009 May 1. DNA Repair (Amst). 2009. PMID: 19410520
-
A DNA glycosylase from Pyrobaculum aerophilum with an 8-oxoguanine binding mode and a noncanonical helix-hairpin-helix structure.Structure. 2005 Jan;13(1):87-98. doi: 10.1016/j.str.2004.10.011. Structure. 2005. PMID: 15642264
-
A distinct TthMutY bifunctional glycosylase that hydrolyzes not only adenine but also thymine opposite 8-oxoguanine in the hyperthermophilic bacterium, Thermus thermophilus.DNA Repair (Amst). 2006 Aug 13;5(8):894-903. doi: 10.1016/j.dnarep.2006.05.001. Epub 2006 Jun 14. DNA Repair (Amst). 2006. PMID: 16781198
-
Biochemical characterization of uracil processing activities in the hyperthermophilic archaeon Pyrobaculum aerophilum.J Biol Chem. 2001 Aug 10;276(32):29979-86. doi: 10.1074/jbc.M102985200. Epub 2001 Jun 8. J Biol Chem. 2001. PMID: 11399761
-
Characterization of a thermostable DNA glycosylase specific for U/G and T/G mismatches from the hyperthermophilic archaeon Pyrobaculum aerophilum.J Bacteriol. 2000 Mar;182(5):1272-9. doi: 10.1128/JB.182.5.1272-1279.2000. J Bacteriol. 2000. PMID: 10671447 Free PMC article.
Cited by
-
Clostridium acetobutylicum 8-oxoguanine DNA glycosylase (Ogg) differs from eukaryotic Oggs with respect to opposite base discrimination.Biochemistry. 2008 Jul 22;47(29):7626-36. doi: 10.1021/bi800162e. Epub 2008 Jun 26. Biochemistry. 2008. PMID: 18578506 Free PMC article.
-
A thermophilic 8-oxoguanine DNA glycosylase from Thermococcus barophilus Ch5 is a new member of AGOG DNA glycosylase family.Acta Biochim Biophys Sin (Shanghai). 2022 Jun 25;54(12):1801-10. doi: 10.3724/abbs.2022072. Online ahead of print. Acta Biochim Biophys Sin (Shanghai). 2022. PMID: 35713316 Free PMC article.
-
Identification of a novel bifunctional uracil DNA glycosylase from Thermococcus barophilus Ch5.Appl Microbiol Biotechnol. 2021 Jul;105(13):5449-5460. doi: 10.1007/s00253-021-11422-8. Epub 2021 Jul 5. Appl Microbiol Biotechnol. 2021. PMID: 34223949
-
Crystal structures of two archaeal 8-oxoguanine DNA glycosylases provide structural insight into guanine/8-oxoguanine distinction.Structure. 2009 May 13;17(5):703-12. doi: 10.1016/j.str.2009.03.007. Structure. 2009. PMID: 19446526 Free PMC article.
-
Structural and functional determinants of the archaeal 8-oxoguanine-DNA glycosylase AGOG for DNA damage recognition and processing.Nucleic Acids Res. 2022 Oct 28;50(19):11072-11092. doi: 10.1093/nar/gkac932. Nucleic Acids Res. 2022. PMID: 36300625 Free PMC article.
References
-
- Scharer O.D. and Jiricny,J. (2001) Recent progress in the biology, chemistry and structural biology of DNA glycosylases. Bioessays, 23, 270–281. - PubMed
-
- Hazra T.K., Izumi,T., Kow,Y.W. and Mitra,S. (2003) The discovery of a new family of mammalian enzymes for repair of oxidatively damaged DNA, and its physiological implications. Carcinogenesis, 24, 155–157. - PubMed
-
- Nash H.M., Bruner,S.D., Scharer,O.D., Kawate,T., Addona,T.A., Spooner,E., Lane,W.S. and Verdine,G.L. (1996) Cloning of a yeast 8-oxoguanine DNA glycosylase reveals the existence of a base-excision DNA-repair protein superfamily. Curr. Biol., 6, 968–980. - PubMed
-
- van der Kemp P.A., Thomas,D., Barbey,R., de Oliveira,R. and Boiteux,S. (1996) Cloning and expression in Escherichia coli of the OGG1 gene of Saccharomyces cerevisiae, which codes for a DNA glycosylase that excises 7,8-dihydro-8-oxoguanine and 2,6-diamino-4-hydroxy-5-N-methylformamidopyrimidine. Proc. Natl Acad. Sci. USA, 93, 5197–5202. - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources

