PROPHECY--a database for high-resolution phenomics
- PMID: 15608218
- PMCID: PMC540080
- DOI: 10.1093/nar/gki126
PROPHECY--a database for high-resolution phenomics
Abstract
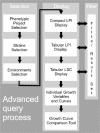
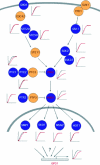
The rapid recent evolution of the field phenomics--the genome-wide study of gene dispensability by quantitative analysis of phenotypes--has resulted in an increasing demand for new data analysis and visualization tools. Following the introduction of a novel approach for precise, genome-wide quantification of gene dispensability in Saccharomyces cerevisiae we here announce a public resource for mining, filtering and visualizing phenotypic data--the PROPHECY database. PROPHECY is designed to allow easy and flexible access to physiologically relevant quantitative data for the growth behaviour of mutant strains in the yeast deletion collection during conditions of environmental challenges. PROPHECY is publicly accessible at http://prophecy.lundberg.gu.se.
Figures


Similar articles
-
PROPHECY--a yeast phenome database, update 2006.Nucleic Acids Res. 2007 Jan;35(Database issue):D463-7. doi: 10.1093/nar/gkl1029. Epub 2006 Dec 5. Nucleic Acids Res. 2007. PMID: 17148481 Free PMC article.
-
High-resolution yeast phenomics resolves different physiological features in the saline response.Proc Natl Acad Sci U S A. 2003 Dec 23;100(26):15724-9. doi: 10.1073/pnas.2435976100. Epub 2003 Dec 15. Proc Natl Acad Sci U S A. 2003. PMID: 14676322 Free PMC article.
-
High-dimensional and large-scale phenotyping of yeast mutants.Proc Natl Acad Sci U S A. 2005 Dec 27;102(52):19015-20. doi: 10.1073/pnas.0509436102. Epub 2005 Dec 19. Proc Natl Acad Sci U S A. 2005. PMID: 16365294 Free PMC article.
-
A picture is worth a thousand words: genomics to phenomics in the yeast Saccharomyces cerevisiae.FEBS Lett. 2009 Jun 5;583(11):1656-61. doi: 10.1016/j.febslet.2009.03.068. Epub 2009 Apr 5. FEBS Lett. 2009. PMID: 19351535 Review.
-
Large-scale mutagenesis and functional genomics in yeast.Funct Integr Genomics. 2002 Sep;2(4-5):193-8. doi: 10.1007/s10142-002-0057-3. Epub 2002 May 9. Funct Integr Genomics. 2002. PMID: 12192592 Review.
Cited by
-
The NPR/Hal family of protein kinases in yeasts: biological role, phylogeny and regulation under environmental challenges.Comput Struct Biotechnol J. 2022 Oct 15;20:5698-5712. doi: 10.1016/j.csbj.2022.10.006. eCollection 2022. Comput Struct Biotechnol J. 2022. PMID: 36320937 Free PMC article. Review.
-
PROPHECY--a yeast phenome database, update 2006.Nucleic Acids Res. 2007 Jan;35(Database issue):D463-7. doi: 10.1093/nar/gkl1029. Epub 2006 Dec 5. Nucleic Acids Res. 2007. PMID: 17148481 Free PMC article.
-
Detecting differential growth of microbial populations with Gaussian process regression.Genome Res. 2017 Feb;27(2):320-333. doi: 10.1101/gr.210286.116. Epub 2016 Nov 18. Genome Res. 2017. PMID: 27864351 Free PMC article.
-
The Molecular Biology Database Collection: 2005 update.Nucleic Acids Res. 2005 Jan 1;33(Database issue):D5-24. doi: 10.1093/nar/gki139. Nucleic Acids Res. 2005. PMID: 15608247 Free PMC article.
-
Accurate, precise modeling of cell proliferation kinetics from time-lapse imaging and automated image analysis of agar yeast culture arrays.BMC Syst Biol. 2007 Jan 8;1:3. doi: 10.1186/1752-0509-1-3. BMC Syst Biol. 2007. PMID: 17408510 Free PMC article.
References
-
- Giaever G., Chu,A.M., Ni,L., Connelly,C., Riles,L., Veronneau,S., Dow,S., Lucau-Danila,A., Anderson,K., Andre,B. et al. (2002) Functional profiling of the Saccharomyces cerevisiae genome. Nature, 418, 387–391. - PubMed
-
- Kamath R.S., Fraser,A.G., Dong,Y., Poulin,G., Durbin,R., Gotta,M., Kanapin,A., Le Bot,N., Moreno,S., Sohrmann,M. et al. (2003) Systematic functional analysis of the Caenorhabditis elegans genome using RNAi. Nature, 421, 231–237. - PubMed
-
- Warringer J. and Blomberg,A. (2003) Automated screening in environmental arrays allows analysis of quantitative phenotypic profiles in Saccharomyces cerevisiae. Yeast, 20, 53–67. - PubMed
MeSH terms
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

