Comparative genome sequencing of Drosophila pseudoobscura: chromosomal, gene, and cis-element evolution
- PMID: 15632085
- PMCID: PMC540289
- DOI: 10.1101/gr.3059305
Comparative genome sequencing of Drosophila pseudoobscura: chromosomal, gene, and cis-element evolution
Abstract
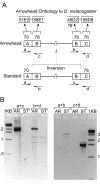
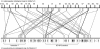
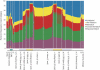
We have sequenced the genome of a second Drosophila species, Drosophila pseudoobscura, and compared this to the genome sequence of Drosophila melanogaster, a primary model organism. Throughout evolution the vast majority of Drosophila genes have remained on the same chromosome arm, but within each arm gene order has been extensively reshuffled, leading to a minimum of 921 syntenic blocks shared between the species. A repetitive sequence is found in the D. pseudoobscura genome at many junctions between adjacent syntenic blocks. Analysis of this novel repetitive element family suggests that recombination between offset elements may have given rise to many paracentric inversions, thereby contributing to the shuffling of gene order in the D. pseudoobscura lineage. Based on sequence similarity and synteny, 10,516 putative orthologs have been identified as a core gene set conserved over 25-55 million years (Myr) since the pseudoobscura/melanogaster divergence. Genes expressed in the testes had higher amino acid sequence divergence than the genome-wide average, consistent with the rapid evolution of sex-specific proteins. Cis-regulatory sequences are more conserved than random and nearby sequences between the species--but the difference is slight, suggesting that the evolution of cis-regulatory elements is flexible. Overall, a pattern of repeat-mediated chromosomal rearrangement, and high coadaptation of both male genes and cis-regulatory sequences emerges as important themes of genome divergence between these species of Drosophila.
Figures

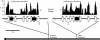


Similar articles
-
Chromosomal rearrangement inferred from comparisons of 12 Drosophila genomes.Genetics. 2008 Jul;179(3):1657-80. doi: 10.1534/genetics.107.086108. Epub 2008 Jul 13. Genetics. 2008. PMID: 18622036 Free PMC article.
-
How malleable is the eukaryotic genome? Extreme rate of chromosomal rearrangement in the genus Drosophila.Genome Res. 2001 Feb;11(2):230-9. doi: 10.1101/gr.162901. Genome Res. 2001. PMID: 11157786 Free PMC article.
-
Rates and patterns of chromosomal evolution in Drosophila pseudoobscura and D. miranda.Genetics. 2006 Jun;173(2):779-91. doi: 10.1534/genetics.105.054585. Epub 2006 Mar 17. Genetics. 2006. PMID: 16547107 Free PMC article.
-
Drosophila as a Model System for Studying of the Evolution and Functional Specialization of the Y Chromosome.Int J Mol Sci. 2022 Apr 10;23(8):4184. doi: 10.3390/ijms23084184. Int J Mol Sci. 2022. PMID: 35457001 Free PMC article. Review.
-
The random versus fragile breakage models of chromosome evolution: a matter of resolution.Mol Genet Genomics. 2007 Nov;278(5):487-91. doi: 10.1007/s00438-007-0287-0. Epub 2007 Sep 13. Mol Genet Genomics. 2007. PMID: 17851692 Review.
Cited by
-
Recombination modulates how selection affects linked sites in Drosophila.PLoS Biol. 2012;10(11):e1001422. doi: 10.1371/journal.pbio.1001422. Epub 2012 Nov 13. PLoS Biol. 2012. PMID: 23152720 Free PMC article.
-
Comparative Expression Dynamics of Intergenic Long Noncoding RNAs in the Genus Drosophila.Genome Biol Evol. 2016 Jun 27;8(6):1839-58. doi: 10.1093/gbe/evw116. Genome Biol Evol. 2016. PMID: 27189981 Free PMC article.
-
Evolutionary Transitions of MicroRNA-Target Pairs.Genome Biol Evol. 2016 Jun 4;8(5):1621-33. doi: 10.1093/gbe/evw092. Genome Biol Evol. 2016. PMID: 27189995 Free PMC article.
-
Specificity and evolvability in eukaryotic protein interaction networks.PLoS Comput Biol. 2007 Feb 16;3(2):e25. doi: 10.1371/journal.pcbi.0030025. Epub 2006 Dec 28. PLoS Comput Biol. 2007. PMID: 17305419 Free PMC article.
-
FISH mapping of microsatellite loci from Drosophila subobscura and its comparison to related species.Chromosome Res. 2010 Feb;18(2):213-26. doi: 10.1007/s10577-010-9112-4. Epub 2010 Mar 3. Chromosome Res. 2010. PMID: 20198419
References
-
- Altschul, S.F., Gish, W., Miller, W., Myers, E.W., and Lipman, D.J. 1990. Basic local alignment search tool. J. Mol. Biol. 215: 403-410. - PubMed
-
- Andersson, B., Wentland, M.A., Ricafrente, J.Y., Liu, W., and Gibbs, R.A. 1996. A “double adaptor” method for improved shotgun library construction. Anal. Biochem. 236: 107-113. - PubMed
-
- Beckenbach, A.T., Wei, Y.W., and Liu, H. 1993. Relationships in the Drosophila obscura species group, inferred from mitochondrial cytochrome oxidase II sequences. Mol. Biol. Evol. 10: 619-634. - PubMed
-
- Bergman, C.M., Pfeiffer, B.D., Rincon-Limas, D.E., Hoskins, R.A., Gnirke, A., Mungall, C.J., Wang, A.M., Kronmiller, B., Pacleb, J.M., Park, S., et al. 2002. Assessing the impact of comparative genomic sequence data on the functional annotation of the Drosophila genome. Genome Biol. 3: RESEARCH0086. - PMC - PubMed
-
- Berman, B.P., Pfeiffer, B.D., Laverty, T.R., Salzberg, S.L., Rubin, G.M., Eisen, M.B., and Celniker, S.E. 2004. Computational identification of developmental enhancers: Conservation and function of transcription factor binding-site clusters in Drosophila melanogaster and Drosophila pseudoobscura. Genome Biol. 5: R61. - PMC - PubMed
Web site references
-
- http://bacpac.chori.org/; BACPAC resources.
-
- http://hgsc.bcm.tmc.edu/projects/drosophila/; BCM-HGSC.
-
- http://pipeline.lbl.gov/pseudo/; VISTA Genome Browser.
Publication types
MeSH terms
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases
