Crystal structure of the virulence gene activator AphA from Vibrio cholerae reveals it is a novel member of the winged helix transcription factor superfamily
- PMID: 15647287
- PMCID: PMC2652724
- DOI: 10.1074/jbc.M413781200
Crystal structure of the virulence gene activator AphA from Vibrio cholerae reveals it is a novel member of the winged helix transcription factor superfamily
Abstract
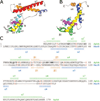
AphA is a member of a new and largely uncharacterized family of transcriptional activators that is required for initiating virulence gene expression in Vibrio cholerae, the causative agent of the frequently fatal epidemic diarrheal disease cholera. AphA activates transcription by an unusual mechanism that appears to involve a direct interaction with the LysR-type regulator AphB at the tcpPH promoter. As a first step toward understanding the molecular basis for tcpPH activation by AphA and AphB, we have determined the crystal structure of AphA to 2.2 angstrom resolution. AphA is a dimer with an N-terminal winged helix DNA binding domain that is architecturally similar to that of the MarR family of transcriptional regulators. Unlike this family, however, AphA has a unique C-terminal antiparallel coiled coil domain that serves as its primary dimerization interface. AphA monomers are highly unstable by themselves and form a linked topology, requiring the protein to partially unfold to form the dimer. The structure of AphA also provides insights into how it cooperates with AphB to activate transcription, most likely by forming a heterotetrameric complex at the tcpPH promoter.
Figures
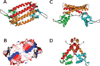

References
Publication types
MeSH terms
Substances
Associated data
- Actions
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

